Surface-enhanced Raman scattering (SERS)—a process that increases the sensitivity of Raman spectroscopy by exploiting surface-plasmon effects—is emerging as a promising nanoscale-imaging modality. Still, while this 30-year-old technique overcomes the sensitivity limitations of standard Raman spectroscopy by increasing the Raman signal up to a millionfold, it suffers from some very practical problems: notably the lack of easily and affordably produced uniform substrates (see www.laserfocusworld.com/articles/274732).
Researchers at the University of Georgia (UGA; Athens, GA) have developed a SERS technique that can detect viruses, and potentially other bioagents, much faster than existing approaches using a method that offers the ability to manufacture consistently uniform silver substrates.1 Their spectroscopic assay, based on SERS using silver-nanorod-array substrates fabricated by oblique-angle deposition, allows for rapid detection of trace levels of viruses with a high degree of sensitivity and specificity, according to Yiping Zhao, assistant professor of physics at UGA.
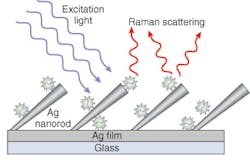
While Raman spectroscopy has previously been used to characterize virus structure, unenhanced Raman spectroscopy has a very small scattering cross section, limiting its use as a low-level bioanalytical sensor. By experimenting with different nanorod structural parameters such as spacing and tilt angle, however, Zhao and chemistry professor Richard Dluhy found a way to significantly amplify the Raman signal. Their patented method involves placing rows of silver nanorods at a density of 13 nanorods/mm2 and a 72°±4° angle from the normal on the substrate that holds the sample (see figure).
“The big problem with SERS is the ability to create a large substrate with high sensitivity that is also very uniform,” Zhao said. “Our technique to fabricate the substrate is thin-film deposition—the same as is used in semiconductor manufacturing—so we can make very uniform nanorods and large-area substrates. These novel substrates allow us to develop SERS-based biosensors rapidly, accurately, and cost-effectively to detect extremely low levels of viruses, thus bridging a critical need for a rapid, sensitive, and reliable means of diagnosing or detecting viruses that does not currently exist.”
Multiple substrates, multiple spots
In initial experiments, the UGA group was able to measure the SERS response of different virus samples and detect differences between viruses, viral strains, and viruses with gene deletions in biological media—all in under a minute. The prototype system comprised a near-IR confocal Raman microscope, a fiber-optic-interfaced 785 nm diode laser, and cooled CCD detectors. The laser power at the sample varied from 10 to 15 mW, with spectral-collection times in the 30- to 50-second range. Using SERS, spectra were collected from multiple substrates and multiple spots across each substrate with enhancement factors of greater than 108. To determine the capacity of SERS to distinguish different RNA viruses, the baseline-corrected enhanced Raman spectra of adenovirus (Ad), rhinovirus (rhino), and human immunodeficiency virus (HIV) have been acquired.
“The results of these studies show that the SERS spectra of viruses can be used to rapidly distinguish between viruses and virus strains, thus serving as a rapid and reproducible means to molecularly fingerprint viruses,” Zhao said. “These results also suggest that it is possible to use SERS to collect the spectra of various viruses and virus strains to develop a reference library of vibrational Raman fingerprints that can be used to rapidly and accurately identify viruses in very small (0.5 to 1.0 µL) volumes.”
Since these initial experiments, Zhao and his colleagues have been working to develop a smaller, less-sensitive, portable system that uses a fiber-optic probe for clinical and field-based applications. Their goal is to enable the technology to be used outside a lab environment for applications such as detecting water pollution, explosives, and nuclear waste.
“This technique saves days to weeks,” said Ralph Tripp, eminent scholar in vaccine development at the UGA College of Veterinary Medicine. “You may someday be able to apply it to a person walking off a plane and know if they’re infected.”
Tripp said the system’s preliminary results are so promising that the researchers are also working to create an online encyclopedia of Raman shift values. With that information, a technician could readily reference a Raman shift for a particular virus to identify an unknown virus.
REFERENCE
1. S. Shanmukh et al., NanoLetters 6 (11) 2630 (2006).