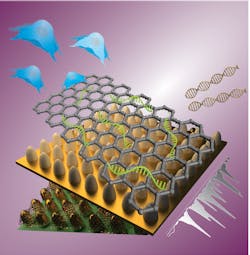
Enhanced Raman biosensor detects stem-cell differentiation
Raman spectral lines, emitted in response to an incident light beam, provide a specific signature—and a powerful method for identifying target molecules. But Raman emission is orders of magnitude weaker than the excitation signal, which limits both sensitivity and the range of applications. An international research team from Rutgers University (New Brunswick and Camden, NJ) and Sogang University (Seoul, South Korea) is developing large-scale nanoarrays that independently modulate both electromagnetic and chemical enhancement mechanisms to improve Raman biosensing of stem cell differentiation (see figure).1
Double-enhanced nanoarray
Very close to a surface (within a nanometer or two), the Raman signal from a given molecule is increased by several orders of magnitude. This surface-enhanced Raman scattering (SERS) is primarily due to a very localized enhancement of the electromagnetic field near the edges of a metallic surface. Steeper edges can enhance the electromagnetic field of an incident light beam. The enhancement depends upon the relationship between the degree of steepness of the metallic surface and the wavelength of light. That is, it depends upon both the composition and roughness of the surface. A surface layered with metallic nanoparticles of varied diameters, for example, will enhance the field of a given wavelength of light to different degrees at different locations. That’s fine for detecting a target molecule, but quantitative measurement requires uniform field enhancement—which, in turn, requires exact control over the surface geometry.
So, the researchers coated a 1 × 1 cm glass surface with photoresist and used the precise control of laser-interference lithography to create polymer features on the substrate. They then used physical vapor deposition (PVD) to deposit a gold layer on the polymer substrate, finding optimum performance with a 40-nm-thick layer. The end result was a regular two-dimensional array of uniform 100-nm-diameter nanocones at a 500 nm pitch. They selected that geometry to maximize the enhancement of a 633 nm incident laser beam.
In addition to the electromagnetic enhancement, surface interactions can also chemically enhance a Raman signal. That is, when a molecule is close enough to a surface, it’s possible that incident light can initiate a charge transfer from the molecule to the surface and back. That resonant charge transfer enhances the Raman signal, if the energy levels of the surface fall in the right place relative to the energy levels of the target molecule.
The Rutgers/Sogang team optimized the chemical enhancement by depositing a 2D graphene oxide (GO) layer on top of the gold. The energy levels of the GO can be tuned by varying the oxidation states of the GO.
The final device is a uniform array of GO-coated gold nanocones, with the electromagnetic enhancement tuned to 633 nm and the chemical enhancement tuned to a specific target molecule. The GO has another convenient property: it forms a strong non-covalent attachment to nucleotide probes. For the proof of principle, the researchers tuned the GO to match the energy level of Cy5 dye.
Biologically relevant sensing
The researchers functionalized the surface with a 21-nucleotide single-strand DNA probe. Each probe was conjugated with a Cy5 dye molecule. The probes adhered to the GO surface, and the nanoarray worked as expected: the Raman signal was enhanced. The nucleotide probes were designed to match a target sequence of interest. When the complementary DNA strand is brought to proximity with the probe, it binds to the probe and releases the Cy5 dye. Theoretically, the Raman signal in a given area, then, is inversely correlated with the concentration of the target DNA sequence.
The researchers calibrated the Raman signal as a function of DNA concentration, where an increase of six orders of magnitude in DNA concentration corresponded to about a factor of five-fold decrease in intensity of a specific spectral peak. The concentration indicated by the nanoarray spectroscopy matched that from polymerase chain reaction (PCR), which is the current gold standard of gene detection. The next step was to apply the method to a clinically relevant problem.
According to Letao Yang, a member of the research team, “The clinical applications of stem cell therapy are limited by concerns about tumor formation or inefficient differentiation.” So the researchers designed nucleotide probe sequences that matched the distinctive markers indicating stem cell differentiation into either neurons or astrocytes. They analyzed a processed extract from differentiated cells. Protein analysis based on immunostaining validated both the selectivity and quantitative accuracy of the SERS nanoarray. Yang noted that this initial success paves the way to “develop an automated biosensing device for the detection of sub-type neural cells,” and possibly “for the detection of vesicles excreted by cells, live cell biosensing, and even sensitive viral detections.”
REFERENCE
1. L. Yang et al., Nano Lett., 19, 11, 8138–8148 (2019).
About the Author
Richard Gaughan
Contributing Writer, BioOptics World
Richard Gaughan is the Owner of Mountain Optical Systems and a contributing writer for BioOptics World.