Light-sheet fluorescence microscopy helps reveal wiring in transparent brains
A team of scientists at the University of Bonn (Germany) and collaborators has harnessed rabies viruses, coupled with a green fluorescent protein, for assessing the connectivity of nerve cell transplants, showing where replacement cells engrafted into mouse brains have connected to the host neural network. Then, a tissue clearing procedure and light-sheet fluorescence microscopy are used to visualize host-graft connections in a whole-brain preparation. The approach opens exciting prospects for predicting and optimizing the ability of neural transplants to functionally integrate into a host nervous system.
Related: Advanced light microscopy enables rapid mapping of brain structure and function in high resolution
Many diseases and injuries result in a loss of nerve cells. Scientists are working on tackling this challenge by transplanting neurons. In Parkinson's disease, for instance, this is attempted with implanted dopamine-producing nerve cells. The key question for such techniques is whether the implanted cells actually connect with the existing neural network of the host brain and thus compensate the functional loss.
“Previous methods only provided an incomplete or very small-scale insight into the functional integration of implanted neurons in the brain,” says Prof. Oliver Brüstle from the Institute of Reconstructive Neurobiology at the University of Bonn and LIFE & BRAIN GmbH (also in Bonn), who led the work.
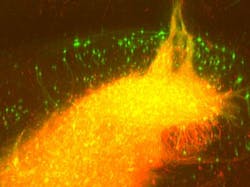
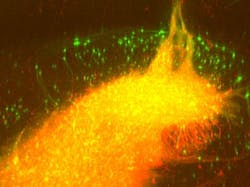
Together with scientists at the Max Planck Institute for Metabolism Research (Cologne, Germany) and the Rosalind Franklin University of Medicine and Science (Chicago, IL), the University of Bonn researchers are exploiting the fact that these viruses spread backwards via synapses. The genetically altered rabies virus, which is no longer dangerous to humans, carries a fluorescent protein. Upon infection of the graft, the transplanted neurons turn green. At the same time, the 'green' virus spreads backwards across established synapses to connected host neurons, which are also turning green.
To visualize the labeled cells, the research team first employed a special tissue clearing procedure, making it possible to turn the brains completely transparent, explains Martin Schwarz from the Bonn Department of Epileptology, who perfected the technique. The transparent brain is then studied layer by layer using a light-sheet fluorescence microscope, which Ulrich Kubitscheck and his team at the Institute for Physical and Theoretical Chemistry at the University of Bonn developed specifically for this purpose.
"With this technique, the brain is scanned in high resolution in over 1000 virtual optical sections—the data is then reconstructed three-dimensionally," Kubitscheck explains. "As the implanted neurons and the recipient's nerve cells connected to them light up green, a three-dimensional brain map can be created that delineates all the recipient cells connected to the transplant—the graft connectome," says Jonas Doerr, who first-authored the study together with Schwarz.
As the brain tissue itself becomes invisible after the clearing procedure, the researchers—in a last step—aligned the fluorescent maps with neuroanatomical data generated via magnetic resonance tomography of mouse brains. "Similar to cities on a globe, all of the cells marked in green can thus be allocated to distinct anatomical territories," says Prof. Mathias Hoehn from the Max Planck Institute for Metabolism Research, whose group conducted the calculations.
"Our findings show that the transplanted neurons integrate in a remarkably region-specific manner into the different transplant sites," Brüstle says.
The researchers hope that the new approach will be particularly useful for studying and optimizing the ability of neuronal transplants to connect with the host brain before they are used for clinical therapy. As a next step, they plan to use the rabies system to investigate how human dopamine-producing cells can be best wired into the brain of mice with induced Parkinson's-like symptoms.
Full details of the work appear in the journal Nature Communications; for more information, please visit http://dx.doi.org/10.1038/ncomms14162.