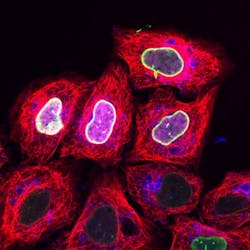
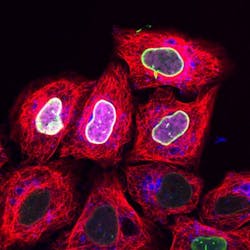
'Cell squeezing' process enables fluorescence microscopy image recording in living cells
Knowing that delivery of protein markers is often insufficient when trying to trace distinct proteins in cells, an international team of researchers from Goethe University (Frankfurt, Germany) and the Massachusetts Institute of Technology (MIT; Cambridge, MA) has found a solution by using pressure to deliver chemical probes in a fine-tuned manner into living cells. In doing so, it allowed them to record fluorescence microscopy images in living cells with high precision and in real time.
Related: Fluorescence microscopy shows how unfolded proteins move in the cell
"Although more and more protein labeling methods utilize synthetic fluorescent dyes, they often suffer from problems such as cell permeability or low labeling efficiency. Moreover, they cannot always be combined with other protein labeling techniques," explains Dr. Ralph Wieneke from the Institute of Biochemistry at the Goethe University.
Recently, the group—led by Wieneke and Prof. Robert Tampé—developed a marker that localizes selected proteins in cells with nanometer precision. This highly specific lock-and-key element consists of the small synthetic molecule trisNTA and a genetically encoded His-tag.
To deliver this protein marker into cells, the researchers used a cell squeezing procedure, in which a mixture of cells together with the marker were forced through narrow constrictions. Under pressure, the cells incorporate the fluorescent probes with an efficiency rate >80%. In addition, the process enabled the researchers to squeeze one million cells per second through the artificial capillary with high throughput.
Since the marker binds efficiently and specifically to the target protein and its concentration can be precisely regulated within the cell, the researchers were able to record high-resolution microscopy images in living cells. What's more, they were able to trace proteins with the marker only when activated by light. Thus, cellular processes can be observed with high precision in terms of space and time.
The researchers can even combine their labeling methods with other protein labeling techniques in living cells to observe several proteins simultaneously in real time. "Utilizing cell squeezing, we were able to deliver a number of fluorescently labeled trisNTAs in cells. This tremendously expands the scopes of conventional as well as high-resolution microscopy in living cells," Tampé explains.
In the future, it will be possible to follow dynamic processes in living cells in time and space at high resolution.
Full details of the work appear in the journal Nature Communications; for more information, please visit http://dx.doi.org/10.1038/ncomms10372.