Protein-based sensor detects viral infection, can kill cancer cells
Bioengineers at the Massachusetts Institute of Technology (MIT; Cambridge, MA) have developed a modular system of proteins that can detect a particular DNA sequence in a cell and then trigger a specific response, such as cell death. The system can be customized to detect any DNA sequence in a mammalian cell and then trigger a desired response, including killing cancer cells or cells infected with a virus, the researchers say.
Related: Disposable photonic sensor detects protein-DNA interactions
James Collins, the Termeer Professor of Medical Engineering and Science in MIT’s Department of Biological Engineering and Institute of Medical Engineering and Science (IMES), is the senior author of a paper describing the work, which is based on a type of DNA-binding proteins known as zinc fingers. These proteins can be designed to recognize any DNA sequence.
“The technologies are out there to engineer proteins to bind to virtually any DNA sequence that you want,” says Shimyn Slomovic, an IMES postdoc and the paper’s lead author. “This is used in many ways, but not so much for detection. We felt that there was a lot of potential in harnessing this designable DNA-binding technology for detection.”
To create their new system, the researchers needed to link zinc fingers’ DNA-binding capability with a consequence—either turning on a fluorescent protein to reveal that the target DNA is present or generating another type of action inside the cell. The researchers achieved this by exploiting a type of protein known as an “intein”—a short protein that can be inserted into a larger protein, splitting it into two pieces. The split protein pieces, known as “exteins,” only become functional once the intein removes itself while rejoining the two halves.
Collins and Slomovic decided to divide an intein in two and then attach each portion to a split extein half and a zinc finger protein. The zinc finger proteins are engineered to recognize adjacent DNA sequences within the targeted gene, so if they both find their sequences, the inteins line up and are then cut out, allowing the extein halves to rejoin and form a functional protein. The extein protein is a transcription factor designed to turn on any gene the researchers want.
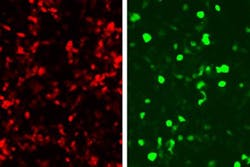
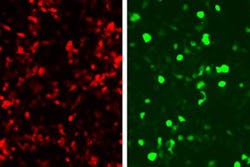
In the study, they linked green fluorescent protein (GFP) production to the zinc fingers’ recognition of a DNA sequence from an adenovirus, so that any cell infected with this virus would glow green. This approach could be used not only to reveal infected cells, but also to kill them. To achieve this, the researchers could program the system to produce proteins that alert immune cells to fight the infection, instead of GFP.
“Since this is modular, you can potentially evoke any response that you want,” Slomovic says. “You could program the cell to kill itself, or to secrete proteins that would allow the immune system to identify it as an enemy cell so the immune system would take care of it.”
The MIT researchers also deployed this system to kill cells by linking detection of the DNA target to production of an enzyme called NTR. This enzyme activates a harmless drug precursor called CB 1954, which the researchers added to the petri dish where the cells were growing. When activated by NTR, CB 1954 kills the cells.
Future versions of the system could be designed to bind to DNA sequences found in cancerous genes and then produce transcription factors that would activate the cells’ own programmed cell death pathways. The researchers are now adapting this system to detect latent HIV proviruses, which remain dormant in some infected cells even after treatment. Learning more about such viruses could help scientists find ways to permanently eliminate them.
While treating diseases using this system is likely many years away, it could be used much sooner as a research tool, Collins says. For example, scientists could use it to test whether genetic material has been successfully delivered to cells that scientists are trying to genetically alter. Cells that did not receive the new gene could be induced to undergo cell death, creating a pure population of the desired cells. It could also be used to study chromosomal inversions and transpositions that occur in cancer cells, or to study the 3D structure of normal chromosomes by testing whether two genes located far from each other on a chromosome fold in such a way that they end up next to each other, the researchers say.
Full details of the work appear in the journal Nature Methods; for more information, please visit http://dx.doi.org/10.1038/nmeth.3585.
Follow us on Twitter, 'like' us on Facebook, connect with us on Google+, and join our group on LinkedIn