CELL BIOLOGY/FLOW CYTOMETRY: Instrumentation advances add flexibility and quantitation to flow cytometry
Fluorescence labels have advanced cell biology and cytometry by enabling the study of cellular function and structure. The ability to attach a wide range of fluorophores to different antibodies has made it possible to classify and map the diverse populations of white blood cells that make up the human immune system, for instance. And the stunning images produced by confocal fluorescence microscopy have not only elucidated the relationships of subcellular structures previously only guessed at, but have also captured the popular imagination (witness the 2014 Nobel Prize in chemistry for super-resolved fluorescence microscopy). Fluorescence-based tools are here to stay.
For all its successes, however, fluorescence tagging seems to have hit a ceiling—even as applications demand ever-greater performance.
One challenge is that fluorophores, the molecules responsible for fluorescent behavior, are not always as well-behaved as we would like them to be. Overwhelmingly organic in nature, their synthesis can be complex, their stability is not guaranteed, and unlike the on-off states of digital switches, their fluorescence performance is governed by absorption and emission spectra-mechanisms firmly rooted in the analog world and subject to nonlinearities. The net effect is considerable uncertainty in the amount of light one can expect a given fluorophore to emit, even with a fixed excitation. When fluorophores are combined—for example, in applications of fluorescence resonance energy transfer (FRET)—the uncertainties add, preventing the results from being reliable quantitatively.
Another problem is that most fluorophores have relatively broad emission spectra—on the order of 30–50 nm. The availability of light sources and detectors (combined with the absorption curve of water) constrains the usable portion of the spectrum to some 400 nm, spanning the visible range and little more. To make the most of this limited range, microscopists and cytometrists have pushed for the development of large libraries of fluorophores with almost any peak emission wavelength in the visible spectrum. But while hundreds of choices exist, only a handful can be used simultaneously, with their spectra spread sufficiently to allow for differential detection. This forces experiments to be run sequentially instead of in parallel—a nuisance in microscopy and a severe limitation in flow cytometry, where sequential interrogation of a single cell is incompatible with the nature of the technique. In other words, the multiplexing capability of fluorescence-based approaches is capped at a low level.
Why fluorescence lifetime?
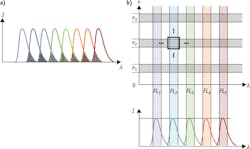
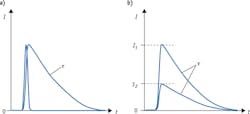
Detection of fluorescence lifetime offers a solution to both problems. When a fluorophore absorbs an incoming excitation photon, it spends a certain amount of time in the excited state before emitting a longer-wavelength photon and returning to its ground state. A collection of identical fluorophores will generate a distribution of time intervals between absorption and emission; this statistical distribution takes the shape of an exponential decay curve (see Fig. 1a), and can be characterized by its 1/e point τ—the lifetime of this fluorescent transition. Critically, the lifetime value τ is completely independent of intensity (see Fig. 1b). This characteristic makes lifetime measurements insensitive to fluctuations in the intensity of the light source, as well as to variations in the amount of light ultimately absorbed by the fluorophores.
By virtue of its insensitivity to intensity fluctuations, fluorescence lifetime lends itself particularly well to quantitative studies. In the case of FRET, the difference between intensity-based and lifetime-based measurements is even more critical, as FRET assays rely on the interaction between two fluorophores. Not only do intensity fluctuations cause uncertainty in the absorption process, but the energy transfer process itself, between the two fluorophores in the FRET pair, is affected by intensity variations—further confounding the results. Measuring fluorescence lifetime makes it possible to eliminate these spurious effects and carry out quantitative FRET applications; for instance, measurements on protein folding and conformation, and on protein-protein interactions.
Fluorescence lifetime has some additional benefits. The lifetime of a fluorescent transition can be influenced by the local molecular environment of the fluorophore. For example, certain fluorophores exhibit a dependence of their lifetime on the pH of the medium surrounding them;1 or they can be sensitive to the concentration of certain ions, such as Ca++.2 This sensitivity can be exploited to use lifetime as a local, molecular-scale probe of environmental conditions.
Worth 1000 pictures
Measurements of fluorescence lifetime have been implemented successfully on imaging platforms. In particular, a technique called time-correlated single-photon counting (TCSPC) has been widely used to generate fluorescence lifetime analysis on advanced microscopes. Developers of TCSPC detection systems, such as Becker & Hickl GmbH and PicoQuant GmbH (both of Berlin, Germany), offer a rich portfolio of products aimed at imaging applications of fluorescence lifetime. The plethora of results3 made possible by their technologies is a testament to the power of fluorescence lifetime imaging microscopy (FLIM) techniques. FLIM-FRET in particular has enjoyed growing popularity.4
However, the slow data acquisition rates of TCSPC, added to the inherent low throughput of microscopy, severely limit the efficiency of analysis. Processing a field of view with perhaps 50 cells can take minutes; while FLIM provides spatially resolved information on each cell, it comes at a steep price in terms of throughput trade-offs. FLIM is a great technique for detailed analysis of handfuls of cells, but it is poorly suited to the rapid analysis or screening of thousands to millions of cells.
For those applications where localization information is not critical, it is highly desirable to trade spatial resolution for analysis throughput. Flow cytometry does just that: By interrogating single cells in a narrow stream flowing at high speeds past a laser interrogation point, it can routinely deliver analysis rates on the order of 10,000 cells per second-three to four orders of magnitude faster than is possible with microscopy-based tools. And while one of the trade-offs is imaging information, flow cytometry offers multiplexed analysis to compensate, in part, for that.
Fluorescence lifetime flow cytometry
It is therefore natural to want to combine the benefits of fluorescence lifetime with the high throughput of flow cytometry. Early work in this direction was spearheaded by John Steinkamp and collaborators at Los Alamos National Laboratory (LANL) in the 1990s,5-8 and briefly explored commercially by BD Biosciences (San Jose, CA) through the work of Robert Hoffman and collaborators9 around the same time. Those undertakings were largely based on analog-electronics implementations of a fluorescence lifetime technique referred to as "phase lifetime" or "frequency-domain lifetime." More recently, researchers at New Mexico State University (NMSU) have revitalized the field with the introduction of new digital fluorescence lifetime technologies for cell sorting and analysis.10 These efforts also involve frequency modulation approaches, and have led to phasor analysis and non-modulated approaches to lifetime.11,12
The early pioneering experiments were crucial for establishing the viability of performing fluorescence lifetime measurements in flow, but traditional instrumentation and signal processing tools used to perform the measurements were complex, expensive, and did not lend themselves to commercialization. For this reason, the new digital approaches are appealing. The frequency-domain approaches, however, can reliably report only a single lifetime component (an area NMSU researchers are now exploring), whereas for the technique to have practical value, flow cytometrists would need it to resolve multiple lifetime components within any single cell.
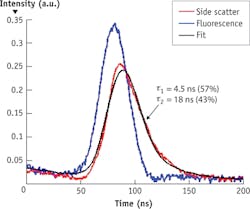
An academic/commercial collaboration (between NMSU and Kinetic River Corp.) supported by the NSF CAREER DBI 1150202 project has produced a new approach that, like its TCSPC FLIM counterpart, is based on the time domain: Fluorescence lifetime values are extracted directly from the fluorescence decay curves emitted in response to pulsed excitation. This has allowed resolution of multiple components of fluorescence lifetime within individual decay curves13—to our knowledge, the first such feat performed at the speeds of particle flow typical of flow cytometry (see Fig. 2).Rapidly growing interest
Other research groups have begun to leverage the unique benefits of fluorescence lifetime coupled with flow cytometry. For example, Miho Suzuki and collaborators of Saitama University in Saitama, Japan, are using FRET bioprobes, pairs of fluorescent proteins and other fluorophores, to study cell signaling for drug discovery applications.14
Figure 4 illustrates one such application-monitoring of protease function. In Fig. 4a, a donor molecule (e.g., GFP or other fluorescent protein) has a site-specific linkage to an acceptor molecule (e.g., fluorescence dyes from the Alexa Fluor, BODIPY, etc. families) using inserted amino acids sequences. The short distance between, and proper alignment of, donor and acceptor ensure that, upon excitation by light in the donor absorption spectrum, efficient FRET takes place, transferring excitation energy to the acceptor molecule. As the donor produces fluorescence emission, the lifetime of the transition is reduced by the presence of the acceptor. Figure 4b shows how, when a protease severs the linker, the donor and acceptor molecules no longer undergo FRET, and the lifetime of the donor emission returns to its longer native value.More broadly, the subject of fluorescence lifetime flow cytometry is enjoying a veritable renaissance. At the last Annual Congress of Cytometry (CYTO, May 2014, Ft. Lauderdale, FL), a new workshop on fluorescence lifetime,15 co-chaired by Silas Leavesley, of South Alabama University (Mobile, AL), featured a distinguished panel (see Fig. 3) that included Prof. Suzuki, who gave the latest results on her work on FRET bioprobes; Patrick Jenkins of NMSU and the Fred Hutchinson Cancer Research Center (Seattle, WA), who talked about using fluorescence lifetime to study cell signaling in yeast cells using FRET; János Szöll"osi of Debrecen University (Hungary), an expert in applying FRET-FLIM to cell signaling pathways in cancer research; and Ralph Jimenez of JILA/NIST and the University of Colorado (Boulder), who presented his work on the relationship of fluorescence lifetime and photostability in fluorescent proteins. Jenkins also demonstrated sorting of cells based on differences in fluorescence lifetime alone.16 And at the most recent Cytometry Development Workshop (October 30–November 2, 2014; La Jolla, CA), fluorescence lifetime flow cytometry was the topic area with the largest number of presented talks, according to the workshop summary by Morgan Richert of the Scintillon Institute.
The future is here
The early interest in multi-exponential fluorescence lifetime flow cytometry has focused on quantitative FRET—a broad set of applications where the ability to accurately resolve multiple fluorescence decay components can make the most immediate difference (see Fig. 4). As researchers become more familiar with fluorescence lifetime, other kinds of applications will follow suit. Researchers at NMSU are pursuing measurements of autofluorescence to differentiate normal from cancerous cells. By measuring the different lifetimes of endogenously fluorescent compound NADH in its free and bound states, one can distinguish cells with normal metabolism and ones with abnormally high metabolic rates—a hallmark of cancer. Being able to do so on a flow cytometry platform would allow rapid discrimination of cancer cells out of large samples of normal background cells. NMSU researchers are also now sorting cells using the fluorescence lifetime as a single parameter.12 Therefore, sorting out cells with altered metabolism based on fluorescence decay times is near.An even broader goal is to use fluorescence lifetime as a separate dimension to massively increase the multiplexing capabilities of flow cytometers (see Fig. 5b). By using fluorescence lifetime as a parameter, spectrally overlapping emissions can be distinguished. Even just doubling the number of channels available for detection—a rather conservative estimate of the potential benefit of multiplexing with fluorescence lifetime—would be a huge boon to immunologists and other cell biology researchers currently forced to work with a limited number of tags on the single cramped dimension of the visible spectrum.
Another tantalizing benefit potentially achievable with fluorescence lifetime multiplexing is the elimination of spectral spillover compensation. By trading some spectral channels for lifetime channels, the remaining spectral channels can be spaced further apart, making compensation unnecessary. As any flow cytometrist will attest, compensation is a source of unending headaches for operators and researchers alike. Eliminating spectral overlap would radically simplify the performance of even complex flow cytometry protocols, speeding up workflow, reducing operating costs, and improving the quality of results. And that's something everyone can get behind!
ACKNOWLEDGEMENTS
The authors recognize the contributions to the Danube I and II projects from Mark Naivar at DarklingX, LLC, with custom data acquisition module design and development and the Kytos data analysis software.
REFERENCES
1. K. M. Hanson et al., Biophys. J., 83, 3, 1682–1690 (Sep. 2002); doi:10.1016/S0006-3495(02)73936-2.
2. K. V. Kuchibhotla, C. R. Lattarulo, B. T. Hyman, and B. J. Bacskai, Science, 323, 5918, 1211–1215 (Feb. 27, 2009); doi:10.1126/science.1169096.
3. W. Becker, "The bh TCSPC Handbook," 5th Edition (Becker & Hickl GmbH, Berlin, October 2012).
4. Y. Sun, C. Rombola, C. Jyothikumar, and A. Periasamy, Cytometry, 83A, 780–793 (2013), and references therein.
5. J. A. Steinkamp and H. A. Crissman, Cytometry, 14, 210–216 (1993).
6. C. Deka, L. A. Sklar, and J. A. Steinkamp, Cytometry, 17, 94–101 (1994).
7. C. Deka et al., Cytometry, 21, 318–328 (1995).
8. C. Deka et al., Cytometry, 25, 271–279 (1996).
9. B. G. Pinsky, J. J. Ladasky, J. R. Lakowicz, K. Berndt, and R. A. Hoffman, Cytometry, 14, 123–125 (1993).
10. J. P. Houston, M. A. Naivar, and J. P. Freyer, Cytometry, 77A, 861–872 (2010).
11. R. Cao, M. A. Naivar, M. Wilder, and J. P. Houston, Cytometry, 85A, 999–1010 (2014).
12. B. Sands et al., PLoS ONE, 9, e109940 (2014); doi:10.1371/journal.pone.0109940.
13. W. Li, G. Vacca, M. Castillo, K. D. Houston, and J. P. Houston, Electrophoresis, 35, 1846–1854 (2014); doi:10.1002/elps.201300618.
14. M. Suzuki et al., Biochim. Biophys. Acta, 1823, 215–226 (2012).
15. J. Houston and S. Leavesley, co-chairs, CYTO 2014: XXIX Congress of the International Society for Advancement of Cytometry, Workshop #7 (Ft. Lauderdale, FL; 2014).
16. P. Jenkins, J. Houston, W. Peria, B. Sands, and R. Brent, CYTO 2014: XXIX Congress of the International Society for Advancement of Cytometry, Paper #97 (Ft. Lauderdale, FL; 2014).
About the Author
Jessica P. Houston
Assistant Professor of Chemical and Materials Engineering, New Mexico State University
Jessica P. Houston, Ph.D., is assistant professor of chemical and materials engineering at New Mexico State University (Las Cruces, NM).
Giacomo Vacca
President, Kinetic River
Giacomo Vacca, Ph.D., earned B.A. and M.A. degrees in physics from Harvard University and a doctorate degree in applied physics from Stanford University. With Nobel Prize winner Bob Laughlin, he developed a novel ultrafast light scattering technique for his dissertation. He has set up entire laboratories from scratch, started and led development programs, and generated intellectual property, with 104 patent applications and 64 patents issued to date. He has also led diverse interdisciplinary groups and managed IP portfolios.
At Abbott Labs, Vacca invented and developed Laser Rastering, a radically innovative concept in flow cytometry that increased the rate of cell analysis by a factor of 30. In 2010 Vacca founded Kinetic River, a biophotonics design and product development company focused on flow cytometry. Since 2017, Kinetic River has been awarded four competitive Small Business Innovative Research (SBIR) grants from the National Institutes of Health, totaling about $2.2 million to date, to help develop innovative flow cytometry technologies. Kinetic River's customers include the National Cancer Institute at the NIH, Italy's National Research Council, and enterprises from startups to Fortune 500 companies.
In 2013 Vacca cofounded BeamWise, a provider of optical system design tools. He is a past Abbott Research Fellow and a senior member of both SPIE and Optica.

![FIGURE 4. Fluorescence lifetime can study apoptosis mediated by proteases. See text for details [12]. FIGURE 4. Fluorescence lifetime can study apoptosis mediated by proteases. See text for details [12].](https://img.laserfocusworld.com/files/base/ebm/lfw/image/2015/12/1501bowvaccaf4.png?auto=format,compress&fit=max&q=45?w=250&width=250)