SINGLE-MOLECULE DETECTION: Detecting single molecules
Microscopy advances continually provide new ways to image and track single molecules within a biological context. Techniques including fluorescence-correlation, super-resolution, and total internal reflection fluorescence (TIRF) microscopy, for instance, provide information on how molecules such as proteins behave in real time in living cells. These forms of imaging are so valuable that super-resolution fluorescence microscopy earned the 2014 Nobel Prize in Chemistry. Other tools, such as photonic microsystems and sophisticated lasers, supply novel ways to trap molecules and illuminate them so that biologists can directly watch them over long periods of time. The best technique, though, depends on the question at hand and the experimental system.
Tracking DNA interactions
At the Laboratory of Nanophotonics and Biosensing at the Max Planck Institute for the Science of Light (Erlangen, Germany), principal investigator Frank Vollmer develops photonic microsystems for single-molecule detection. He says, "Light is good at creating photonic biosensors that are ultrasensitive, and we can detect things without labels, meaning that molecules aren't modified by anything."
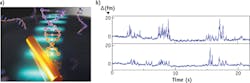
From a simplified perspective, a photonic microsystem resembles two mirrors facing each other so that light can bounce back and forth between them. "You can confine the light in microstructures," says Vollmer. "These devices could be microcavities on a microchip where light interacts many thousands of times with biomolecules." Such a large number of interactions allows single-molecule detection without any label (see Fig. 1).
To demonstrate this technology, Vollmer monitored single DNA molecules on his microdevice, as explained in his 2014 article in Nature Nanotechnology. "It is difficult to detect single DNA fragments in the optical domain without labels," he says. "That had not been done before." Vollmer and his colleagues attach a single-stranded DNA 'bait' to their microdevice, which they then immerse in liquid. When a matching—or complementary—DNA fragment binds from solution to the bait, these single-molecule contacts can be tracked over time. So it is possible to investigate how long the DNA fragments interact with each other and how often the bait captures a segment. "This approach makes it possible to use a single DNA receptor and to follow its successive interactions with various DNA segments in the sample solution," says Vollmer. "Based on the duration and frequency of the measured contacts, it is then possible to detect specific unlabeled DNA molecules."
Resolving RNA
Some of the future's biggest biological discoveries could come from RNA. Until now, many questions about this nucleic acid remain unanswered. As Nils G. Walter, professor of chemistry and director of the Single Molecule Analysis in Real-Time (SMART) Center at the University of Michigan at Ann Arbor, says, "About 75 percent of the human genome codes for RNA, compared to just under two percent for protein, and we are only beginning to understand what all of that RNA is doing."
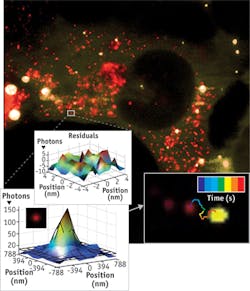
As part of Walter's research, he explores RNA at the single-molecule level. For example, he and his team use a TIRF microscope to manipulate cells in culture (see Fig. 2). "You can inject the RNA of interest with a fluorophore label attached and then track the RNA. You can see if and where it diffuses and see if that changes over time," Walter says. To combine single-molecule information, Walter also employs computational approaches.Studying exocytosis
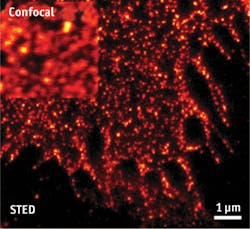
Single-molecule detection depends on sophisticated systems that in turn depend on capable components. As examples, Håkan Karlsson, CEO of Cobolt (Solna, Sweden), points to fluorescence-correlation microscopy and super-resolution microscopy, which need high-performance lasers offering "low intensity noise and very good beam quality." Fluorescence correlation has benefited from lasers at several wavelengths, but mostly in the turquoise and green ranges, while higher-power lasers have enabled super-resolution microscopy. As an example, Karlsson mentions that the Cobolt Flamenco 660 nm single-frequency diode-pumped solid-state (DPSS) laser works very well for stimulated emission depletion (STED) microscopy, offering a significant reduction in size and complexity of the setup. In addition, its low-intensity noise enabled the combination of STED and fluorescence-correlation spectroscopy to study diffusion processes of lipids and proteins at nanoscale levels in the cellular plasma membrane (see Fig. 3).Sebastian Barg, associate professor in the department of medical cell biology at Uppsala University in Sweden, uses such lasers in a home-built TIRF microscope, created from a Zeiss platform used as the base. Compact size is "very, very good," says Barg, adding that "These lasers don't take much power, and they last a long time."
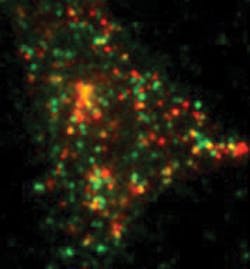
With this system, Barg tracks single proteins that have been tagged with a version of green or red fluorescent protein—GFP or RFP, respectively (see Fig. 4). For example, he studies the process of exocytosis, in which hormone- or neurotransmitter-filled vesicles get released into the blood stream or at connections between neurons. Traditionally, scientists suspected that the molecular machinery that fuses vesicles to the membrane for exocytosis is always in place. By watching the behavior of single molecules, though, Barg showed that the machinery for exocytosis gets assembled when a vesicle touches the membrane. "This process involves at least 30 different proteins and some lipids, too," Barg says. "To see this, you need to look at single molecules within the ensemble."Broader insight
Many of the single-molecule approaches will produce data and information that scientists will use to understand bigger systems, such as a living cell. As Walter says, "We're trying to understand emergent behavior for larger systems from component behavior." He adds, "Life emerges from the stochastic behavior of single molecules in aggregate." Today's work on single-molecule detection promises to—molecule by molecule—help biomedical scientists and clinicians build an ultimate understanding of cellular life.
About the Author
Mike May
Contributing Editor, BioOptics World
Mike May writes about instrumentation design and application for BioOptics World. He earned his Ph.D. in neurobiology and behavior from Cornell University and is a member of Sigma Xi: The Scientific Research Society. He has written two books and scores of articles in the field of biomedicine.