Color-coded quantum dots provide fast DNA testing
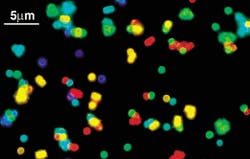
Indiana University (Bloomington, IN) researchers have developed a multicolor optical-coding method that identifies tens of thousands of genes all at once by using tiny semiconductor crystals that dazzle in ultraviolet light (see Fig. 1). The technique works like a bar code—with each color and intensity combination corresponding to an individual gene—and can study up to 40,000 genes or proteins in as little as 10 minutes.
Researchers have tried for many years to use tiny crystals, called quantum dots, as glowing labels for genes, proteins, and other biological molecules. Quantum dots promise to be faster in binding kinetics, more flexible in target selection, and less expensive as tests for on-the-spot biological analysis or patient diagnosis. Until now, they have been difficult to collect and manipulate with enough precision to be useful.
"We solved all of the technical problems," said biomedical engineer Shuming Nie, who led the research effort. "The idea is very simple and straightforward, but I think we're the first ones to make it work."
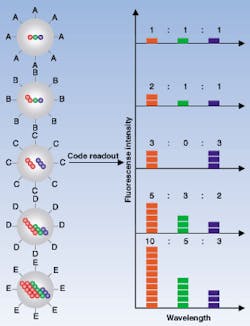
Nie's research team discovered a way to capture the quantum dots in specific quantities and in a wide range of colors and various intensities. The researchers made porous microbeads of polystyrene as small as 1 nm in diameter and seeded them with zinc sulfide-capped (ZnS) cadmium selenide (CdSe) nanocrystals (see Fig. 2). Both beads and quantum dots were made water-repellent to encourage the quantum dots to move into the pores. Once the quantum dots infiltrated the pores, they were sealed in. In ultraviolet light, each dot radiates a brilliant color. Novel optical properties, such as size-tunable emission and simultaneous excitation, also make the highly luminescent quantum dots ideal fluorophores for wavelength-and-intensity multiplexing.To demonstrate the use of these quantum dots in DNA analysis, the researchers prepared microbeads of three colors, or spectral wavelengths, and attached them to strips of genetic material. Each color corresponded with a specific DNA sequence. These were used as probes to seek out complementary pieces of genetic material in a DNA mixture. The results confirmed that coding and target signals can be simultaneously read at the single-bead level.
Imaging and spectroscopic measurements indicate that the quantum dot-tagged beads are highly uniform and reproducible, yielding bead identification accuracies as high as 99.99% under favorable conditions. The researchers believe that this spectral coding technology will open new opportunities in gene expression studies, high-throughput screening, and medical diagnostics.
REFERENCE
- M. Han et al., Nature 19, 7 (2001).
About the Author
Sally Cole Johnson
Editor in Chief
Sally Cole Johnson, Laser Focus World’s editor in chief, is a science and technology journalist who specializes in physics and semiconductors.