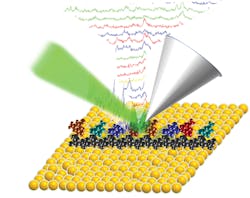
TERS images single-stranded DNA without amplification
Optical imaging at the single-molecule level is often carried out in a low-temperature environment to minimize thermal sensor noise. To achieve high sensitivity as well as ultrahigh spatial resolution in ambient conditions, a joint research group from Texas A&M University (College Station, TX), Rensselaer Polytechnic Institute (RPI; Troy, NY), Friedrich-Schiller-Universität Jena (Germany), and Baylor University (Waco, TX) used tip-enhanced Raman spectroscopy (TERS) to sequence a single-stranded DNA molecule (from the M13mp18 bacteriophage) with an accuracy of more than 90% and a resolution of 0.5 nm.
The TERS method relies on surface plasmon effects in noble-metal probe tips. In the presence of the electromagnetic field, when the plasmon is resonant with the excitation wavelength, intensities of the localized electromagnetic field near the tip are greatly enhanced, increasing Raman scattering and overcoming the diffraction limit.
In this study, the researchers used gap-mode TERS in which the tip and a nearby gold film form a plasmonic nanoscale gap, allowing for nanoscale sensing and single-molecule detection based on strongly enhanced Raman signals. The TERS Raman spectra were used to identify the individual DNA nucleotides through careful spectral analysis with correlation coefficients. By resolving single molecules, TERS imaging of DNA can be used to examine its spatial alignment, structure, and sequence in a timely manner (four seconds per base) without the need for fluorescent labeling or amplification. Furthermore, this technology is likely applicable for scanning RNA, providing a route for easy and economical direct RNA sequencing without making complementary DNA (cDNA) or involving polymerase chain reaction (PCR) amplification, which, taken together, could supplant RNA sequencing platforms. Reference: Z. He et al., J. Am. Chem. Soc., 141, 753–757 (2019).
About the Author

Gail Overton
Senior Editor (2004-2020)
Gail has more than 30 years of engineering, marketing, product management, and editorial experience in the photonics and optical communications industry. Before joining the staff at Laser Focus World in 2004, she held many product management and product marketing roles in the fiber-optics industry, most notably at Hughes (El Segundo, CA), GTE Labs (Waltham, MA), Corning (Corning, NY), Photon Kinetics (Beaverton, OR), and Newport Corporation (Irvine, CA). During her marketing career, Gail published articles in WDM Solutions and Sensors magazine and traveled internationally to conduct product and sales training. Gail received her BS degree in physics, with an emphasis in optics, from San Diego State University in San Diego, CA in May 1986.