Second-harmonic-generation microscopy provides clues to brain-cell development
The generation of ultraviolet (UV) light using second-harmonic generation (SHG) can be exploited in microscopy, in which a specimen can emit SHG light that clarifies certain structures of interest in the specimen. Given its capability for detecting signals in collagen, SHG microscopy is finding favor in biology and medicine, particularly for research and diagnostic applications in the brain.
A research collaboration between biophysicists from Cornell University (Ithaca, NY) and Harvard University (Cambridge, MA) is using SHG imaging to highlight the cytoskeletal infrastructure of nerve cells and map the nervous system as it develops and struggles to repair itself. The research is expected to find application in the study of Alzheimer's and other neurodegenerative diseases, and possibly in helping develop regrowth techniques for patients with spinal-cord injuries.
Under the direction of Watt Webb, professor of applied physics at Cornell, the research group is focusing on detecting polarity in microtubule assemblies in living brain tissue. In developing nerve cells, microtubules are extensions from the cell body that grow to form dendrites and axons. Microtubules are made of tiny polymers and are a major part of the cellular cytoskeleton, responsible for mechanical support and intracellular transport. The researchers say that SHG imaging capitalizes on a structural polarity that exists in the polymers, making the characteristics at one end different from the other. Changes in this polarity are key to how neurons grow and find their orientation in the developing brain.
"Never before has there been a satisfactory way of detecting polarity in microtubule assemblies in living brain tissue," Webb said. "Now we can follow the development of microtubules in vivo to see how architectural changes are occurring in nerve cells or in any other living cells where microtubules are found."
Researchers have attempted similar work using electron microscopy and culture dishes, but that approach requires the cell sample to be fixed, which means the sample is killed and thus can only provide information about the cell in a certain time and place.
Demonstration
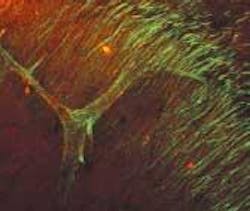
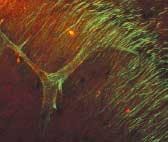
To demonstrate the SHG technique, Webb's group depicted axon bundles and individual axons in rat hippocampal brain tissue and axons growing from cell bodies in culture dishes (see figure). Second-harmonic generation and two-photon-fluorescence (TPF) microscopy were simultaneously performed on a modified inverted Olympus IX-70 microscope. The excitation source was a modelocked diode-pumped Ti:sapphire laser (100-fs pulses at 80 MHz). The laser polarization was controlled via a Berek polarization compensator and the beam focused into the sample. The resulting SHG emission was collected in a forward-propagating direction, while the TPF was epi-collected through the excitation objective. A combination of dichroic mirrors, bandpass and blue-glass filters, and polarization analyzers separated the signals for detection with photomultiplier tubes.
The SHG spectra were recorded by stage scanning the sample and coupling the transmitted light into a liquid-nitrogen-cooled fiber-coupled spectrometer. Relative effective SHG cross sections were corrected for transmission of optics, power at the sample, pulse width, and so on. Propagation direction distributions were deduced from images of SHG from so-called "mossy" fibers in the brain cells in both forward and epi-propagating directions with fluorescence calibration for absorption and scattering in the tissue and collection efficiency differences of the optics and detectors in each direction. The signal from a single microtubule is not enough to detect, so the uniform polarity is key to enabling the SHG signal to be detected.
"Microtubules with uniform polarity generate a second harmonic, but microtubules with mixed polarity don't," said Daniel Dombeck, a graduate student in the School of Applied and Engineering Physics at Cornell and a member of Webb's research team. "We get destructive interference instead so that the axons light up and dendrites and everything else with nonuniform polarity in the microtubules stay dark."
According to Dombeck, microtubules—which have their own inherent spatial polarity—are important indicators of what neurons are doing at a certain time. When a neuron begins to develop, it starts out as a little ball and then sends out processes that become the dendrites and axons. But how neurons know how to do this and how they know how to define their polarity in a certain way remains a big question in neurobiology. In addition to helping decipher how the brain becomes "wired," SHG imaging may also reveal the relationship between microtubule polarity and an overexpression of the tau protein, which is found in the brain tissue of Alzheimer's patients.
In other experiments involving non-neuronal structures, the researchers were able to image microtubules in the mitotic spindles of dividing cells and microtubule-based cilia that line the inner walls of the aquaductus cerbri and "waggle" to propel fluid through the brainstem. When individual successive images are assembled into a video, cell division can be followed and the fluid-propelling motion of brainstem duct cilia studied in detail.