OPTOELECTRONIC APPLICATIONS: BIOANALYTICS: Plasmons point out proteins

Life depends on proteins. These molecules drive reactions, make up receptors that participate in diseases, and much more. Many biochemical processes, though, depend on multiple proteins, often interacting in a cascade. But determining and measuring which protein interacts with which, and to what degree, is difficult at best. A researcher might label one protein with a green fluorescent dye and another with a red fluorescent dye; if the two labels appear on top of one another it can be assumed that these proteins interact, but the labels themselves can change how proteins behave. Scientists in biochemical research, such as those in drug discovery, need to see interactions of proteins in their natural form, and plasmon resonance allows them to do that.
A plasmon is a collective oscillation of electrons at the interface of a metal and a dielectric (see www.laserfocusworld.com/article/279840). Triggering that oscillation requires radiation of a specific wavelength, which depends on the kind of electrons involved and nearby surfaces. Even proteins attaching to a surface change the frequency at which resonance occurs. Measuring that resonance, especially its changes, tells a researcher something about what’s going on at a surface, such as one protein binding to another. And that’s just the tip of this technological iceberg. When it comes to protein-protein interactions, plasmon resonance reveals even more.
“You can get the thermodynamic binding constant and kinetic data,” said Katherine Willets, assistant professor of chemistry at the University of Texas (Austin, TX). “In terms of how a biological system works, that knowledge is really helpful.”In fact, protein interaction arrays may well prove to be the link between the vast repository of data that has emerged from proteomics initiatives and the world of clinical and biological research.
Willets uses localized surface plasmon resonance (LSPR), in which a plasmon occurs on a metallic nanoparticle smaller than the wavelength of visible light. These devices sense the binding of molecules that exist in as little as zeptomolar concentrations-just 10-21 molar. In addition, these nanoparticles only measure nearby molecules, those within roughly 5 nm. Even better, LSPR nanoparticles can be fixed to a glass substrate in an array and “measured” by eye because the array starts as one color and ends up a different color. In addition, the color change can be measured with a miniature spectrometer.
Instead of using nanoparticles, a metallic film on a chip can generate surface plasmon resonance (SPR) when illuminated. These sensors cover more ground, detecting molecules within 200 nm of the surface. In both cases, the metallic component can be functionalized. That is, a ligand-say, an antibody-can be attached, which creates target specificity, according to Willets. That ligand picks out specific molecules in a sample, and a plasmon-resonance measurement keeps track of the binding.Getting started
In 1990, Ulf Jönsson and his colleagues at Pharmacia Biosensor (later Biacore, now part of GE Healthcare) used SPR as the engine in a biosensor. Today’s Biacore sensor consists of a chip covered with a gold film that can be studded with molecules that capture peptides, proteins, nucleic acids, or carbohydrates. A microfluidic system then flows a sample over the chip, and a wedge of polarized light-created from a near-infrared LED, the peak wavelength of which depends on the specific system-illuminates it from a range of angles. This light triggers SPR, which dims the reflected light. A CCD camera or diode array can detect the resonance angle (the angle of incidence that reduces the reflected light the most). That angle depends on the refractive index at the chip-sample interface, and it reveals the molecular mass-or how much target is bound. Different binding molecules can even be arranged in an array on one chip to simultaneously measure the interactions with hundreds of targets. A successful instrument, though, depends on more than the parts.
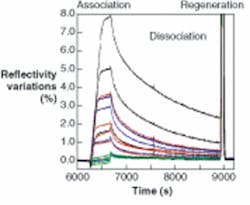
“We are focusing on developing whole-instrument systems for life-science customers,” said Stefan Löfås, chief scientific officer at Biacore International (Uppsala, Sweden; see Fig. 1). “The optical sensor is just one part of the total system.” The optical sensor produces a sensorgram. As binding occurs, Löfås said, the response increases to a maximum, and then falls off as molecules dissociate (see Fig. 2).
To make this all run smoothly, the parts must all work together, he added. The optics detection is integrated with fluidics and modified surfaces, plus software for running the instruments.
Spawning systems
Other companies are also building SPR systems. GenOptics (Orsay Cedex, France), for example, makes its SPRi-Biochip to run in its SPRi-Plex or SPRi-Lab+ systems (see Fig. 3). This chip can be spotted with as many as 400 binding molecules, including antibodies, proteins, peptides, DNA, and sugars, according to Philippe Kerourédan, CEO of GenOptics. He notes, however, that peptides often work better than complete proteins.“Proteins need the right three-dimensional shape to be active, and when you attach a protein to the surface of a biochip you risk damaging the activity,” he said.
The GenOptics systems use LED illumination at about 810 nm and capture the reflection with a CCD camera-a setup Kerourédan calls “nothing special.”
“The SPR principle is well known, but its implementation requires a high degree of know-how,” he said. “The innovation is the way you organize all of these things. GenOptics has protected its way of assembling the optical parts as well as the chemistry to attach the biomolecules onto the biochip.” In addition, because no labels are needed, the proteins keep their natural conformation.
An SPR chip can also be labeled with proteins related to a specific disease. Using a certain allergy, for example, a researcher would attach antigens to an SPR sensor and regularly run a patient’s sera over the allergen-studded chip to see if the patient’s antibodies react with the antigens.
“This should give the clinician valuable information on the relevance of the antibodies produced by a single patient,” Kerourédan said. “Allergens recognized with a high affinity might be more clinically relevant than others, which would be recognized with only low-affinity antibodies. This information should have potential predictive values on the evolution of the disease or the effects of vaccination on the production of protective antibodies.”
SPR-based sensors also have potential use outside the pharmaceutical industry. GenOptics, for example, has been collaborating with the French army to develop sensors that detect different pathogens that could be disseminated by terrorists. “You could get a specific reading of what’s been released, all in just a few minutes, but this kind of application is not yet routine,” Kerourédan said.
Aiming at antibodies
Instead of creating a system that detects many types of molecules, Plexera (Bothell, WA) targeted the specific needs of the pharmaceutical industry by visiting more than 40 companies and running focus groups. The result? Antibodies-lots of antibodies. Thus, Plexera’s NanoCapture Microarray provides room for 5000 spots, which could mean 5000 different antibodies on one chip. A customer uses a protein printer-validated by Plexera-to spot a microarray with complete antibodies or pieces of them. Plexera’s ProteomicProcessor reads the results. Even with 5000 antibodies on a microarray, a scientist can analyze the binding of one antigen in just 10 minutes.
“If you want definitive kinetic data from high-affinity antibodies with a slow off rate, it could take hours, as with any SPR system,” said Ron Dudek, director of business development at Plexers. “We believe this type of throughput will greatly increase the efficiency and productivity of the drug-discovery process.”
Antibody sensors can also be focused on specific tasks. Plexera is already collaborating with scientists at the Institute of Systems Biology (Seattle, WA) to find biomarkers of liver toxicity. Then, an SPR sensor can be tagged with antibodies that pick up these biomarkers. Pharmaceutical companies could use such a sensor to find out-possibly very early in development-whether a new drug could damage a person’s liver.
Pushing plasmons ahead
Sensors based on plasmon resonance continue to cover more territory in the biological sciences. For example, Biacore’s instruments can already sense very small compounds, including pieces of drugs called fragments. Many pharmaceutical companies now test fragments for binding to a disease target, and then combine several of them into a potential new drug. Biacore’s systems can test a thousand fragments a day for binding strength.
LSPR also shows biomedical potential. Richard Van Duyne of Northwestern University developed a sensor based on a mica substrate, then a layer of silver nanoparticles, followed by a self-assembled monolayer, and, finally, antibodies for amyloid beta proteins, which are associated with Alzheimer’s disease (see www.laserfocusworld.com/article/229675). Then, by tracking LSPR with this sensor exposed to Alzheimer-related proteins, Van Duyne could distinguish the concentration of these proteins in samples, such that they might be measured as biomarkers for this disease.
Many more advances lie ahead for plasmon resonance in the life sciences. For one thing, scientists would like to take plasmon-resonance sensing to the biology, instead of bringing the biology to the sensor. Nanoparticles could help with that, but sensing LSPR from disparate locations-say, throughout an animal’s body-remains challenging. In the meantime, many scientists continue to count on oscillations of electrons to probe biology from many angles.
About the Author
Mike May
Contributing Editor, BioOptics World
Mike May writes about instrumentation design and application for BioOptics World. He earned his Ph.D. in neurobiology and behavior from Cornell University and is a member of Sigma Xi: The Scientific Research Society. He has written two books and scores of articles in the field of biomedicine.