Advances in Imaging: 3D superresolution imaging and tracking using a double-helix point-spread function
ANURAG AGRAWAL and LESLIE KIMERLING
Optical microscopes are indispensable tools in numerous fields, including biology, medicine, and materials science. Traditionally only used for qualitative observation, they are now extensively used to obtain quantitative data about the specimen. Often, this involves imaging of cell structures or accurately tracking discrete particles in space and time, also known as single-particle tracking (SPT). However, the achievable resolution of traditional light microscopy is limited by the diffraction of light to approximately 200 nm in the lateral dimensions (x-y) and 500 nm in the axial dimension (z).
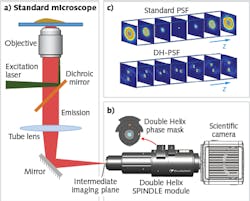
The recent advent of superresolution imaging has enabled researchers to surpass the diffraction barrier and visualize subcellular structures well below the 200 nm limit.1 Although the highest-resolution methods are able to achieve imaging precision on the order of 10 to 20 nm laterally, they typically lack high axial resolution and range. Localization imaging methods work by fitting the image of a point emitter to a known analytical function resembling the point-spread function (PSF) of the microscope, which in the case of a standard microscope resembles an hourglass shape when traced in 3D. A new class of engineered PSFs enables imaging with both high axial precision and an extended axial depth. The 3D trace of a family of these engineered PSFs resembles two intertwined helices and are collectively known as the double-helix (DH) PSF (DH-PSF).2
The DH-PSF is typically generated by placing a phase plate mask in the detection path of the optical imaging system. An implementation of this technique is currently available in the form of an optical module—the SPINDLE—that can be easily integrated with most scientific microscopes and cameras (see Fig. 1). As shown in Fig. 1c, by using the DH-PSF, the image of each point emitter is rendered in the form of two lobes. The center of the pair of lobes determines the lateral position of the emitter and the angle between the lobes determines the axial position.
This additional information facilitates superresolution reconstruction at very high precision (<30 nm) in the lateral and axial dimensions for the typical low photon numbers emitted by individual molecules. Additionally, and importantly, the DH-PSF also extends the depth of field over which molecules can be localized to up to 10 µm. The DH-PSF, in conjunction with matched algorithms, extends the imaging capabilities of a microscope to nanometer-scale 3D localization for imaging3, 4 and tracking.5, 6 To elucidate this further, three case studies are presented.Case study: Capturing information otherwise lost
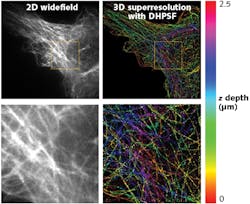
In this study, microtubules in a mouse embryonic fibroblast (MEF) were observed with the SPINDLE, allowing 3D superresolution images to be reconstructed. The DH-PSF 3D superresolution reconstructions here are shown compared to standard widefield fluorescence microscopy images in Fig. 2.As shown in Fig. 2, the 3D DH-PSF superresolution reconstruction captured a depth of 2.5 μm. When compared to the 2D widefield image, the DH-PSF 3D images show precise reconstructions of microtubules that were otherwise lost in the background (see Fig. 2; bottom images). Moreover, the extended depth of field of the DH-PSF enabled a fuller imaging and reconstruction of the microtubules for easy distinction of microtubules at different z depths of the cell. This extended depth range has opened a plethora of opportunity for extended study of biology, from immunology to dementia.
Case study: Probing the structure of stress granules
An enigmatic pathological hallmark of numerous neurodegenerative diseases is the accumulation of potentially toxic RNA and protein aggregates called stress granules. At the University of Colorado, Boulder, Roy Parker’s lab is interrogating the properties of stress granules and how these properties may become altered in neurodegenerative disease.
Stress granules arise in the cytoplasm after a cell is exposed to stress such as oxidative stress. Altered stress-granule regulation may lead to the aberrant accumulation of potentially toxic RNA-protein stress-granule-like aggregates. Conventional light microscopy reveals stress granules as loosely structured aggregates composed of the RNA-binding protein G3BP1. Using confocal microscopy and genetic engineering, the Parker lab fluorescently tagged G3BP1 and confirmed that G3BP1 is an essential scaffolding component of stress granules. To better understand how G3BP1 contributes to the scaffolding of stress granules, the Parker lab turned to using the DH-PSF in conjunction with superresolution imaging. Using the DH-PSF and superresolution imaging, the Parker lab was able to image stress granules with sub-diffraction limited precision in all three dimensions.4
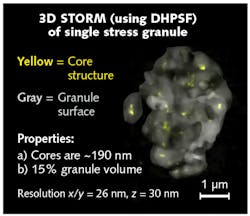
Superresolution imaging revealed, for the first time, that the underlying structure of stress granules is made, in part, of core G3BP1 substructures (see Fig. 3). Armed with this new insight on the substructure of stress granules, the Parker lab is now exploring how stress-granule substructure may become altered in neurodegenerative diseases, opening new avenues for developing therapeutic strategies for targeting these stress granules in disease.Case study: Understanding interfacial hopping of molecules
Trafficking of nutrients into the cell, the signaling of information to the cell nucleus, and the initiation of cell division are all fundamental areas of the study of cellular dynamics, important to a range of biomedical areas of study. The Schwartz lab of the University of Colorado, Boulder studies the dynamic behavior of molecules and nanoparticles in confined environments using fluorescence microscopy in combination with 2D single-particle tracking. However, further investigations have shown that to completely understand interfacial transport, it is important to understand the 3D motion of the molecules during their 3D flights and the surface capture process.
Using DH-PSF technology in conjunction with the SPINDLE, the researchers were able to extend their multiparticle tracking to three dimensions. In a recently published journal article,6 they tagged individual human serum albumin molecules with a fluorescent dye, capturing the trajectories of individual molecules moving along negatively charged silica surfaces (see Fig. 4).The team mapped the 3D motion of a molecule with a spatial precision of 20 nm in the axial direction and 15 nm in lateral directions while maintaining a temporal precision of around 0.1 s. By modifying the surfaces with various silane compounds, the team observed, for the first time, fast 3D molecular hops and, in some instances, “flights” through the adjacent liquid phase followed by recapture at the solid surface.
Since the publication of the work in the journal Physical Review Letters, a viewpoint article was subsequently published.7 The authors of the viewpoint article, Bar-Ilan University physicists Eli Barkai and Yuval Garini, identified the experiments as an “important contribution of interactions between molecules and surfaces in determining the dynamics of the molecules, something that is not yet fully understood,” providing a more detailed picture of the behavior of proteins and other biomolecules in cellular processes. This work is not only important to areas of biomedicine, but is also valuable to multiple industrial applications, including pharmaceutical purification and wastewater remediation.
ACKNOWLEDGEMENT
SPINDLE is a trademark of Double Helix.
REFERENCES
1. E. Betzig et al., Science, 313, 5793, 1642–1645 (Sep. 2006).
2. S. R. P. Pavani and R. Piestun, Opt. Express, 16, 5, 3484 (Mar. 2008).
3. S. R. P. Pavani et al., Proc. Natl. Acad. Sci. USA, 106, 9, 2995–2999 (Mar. 2009).
4. S. Jain et al., Cell, 164, 3, 487–498 (2016).
5. S. R. P. Pavani and R. Piestun, Opt. Express, 16, 26, 22048 (Dec. 2008).
6. D. Wang, H. Wu, and D. K. Schwartz, Phys. Rev. Lett., 119, 26, 268001 (Dec. 2017).
7. E. Barkai and Y. Garini, “Viewpoint: 3D imaging of hopping molecules,” Physics, 10, 139 (Dec. 2017).
Anurag Agrawal is principal optics scientist and Leslie Kimerling is co-founder and CEO, both at Double Helix, Boulder, CO; e-mail: [email protected]; www.doublehelixoptics.com.
![FIGURE 4. Representative 2D (a) and DH-PSF 3D (b) trajectories are shown for human serum albumin on an amino-silane modified fused silica surface; the inset shows the corresponding trace of z position vs. time [6]. FIGURE 4. Representative 2D (a) and DH-PSF 3D (b) trajectories are shown for human serum albumin on an amino-silane modified fused silica surface; the inset shows the corresponding trace of z position vs. time [6].](https://img.laserfocusworld.com/files/base/ebm/lfw/image/2018/05/1804lfw_kim_f4.png?auto=format,compress&fit=max&q=45?w=250&width=250)