Defocused microscope reveals macrophage antics
By defocusing an optical microscope, researchers at the Federal University of Minas Gerais (Belo Horizonte, Brazil) have reported a simpler and more direct method of analyzing the behavior of scavenging white blood cells than they say would have been possible using phase-contrast microscopy.1
The phase-contrast method is useful for viewing phase objects, such as cellular structures, because the illumination of a nonuniformly transparent specimen causes diffraction-induced phase changes in the transiting light that show up in the resulting image as proportional changes in intensity. So phase-contrast microscopy can measure the thickness of a thin phase object with a uniform index of refraction.
Oscar Mesquita and his colleagues were primarily interested in observing dynamics of cell surface curvature, however, to better understand how scavenging white blood cells (macrophages) go about consuming their prey is a process called phagocytosis. They observed that the surface curvature of a thin phase object with a uniform index of refraction was actually proportional to the image contrast obtained with a defocused optical microscope.
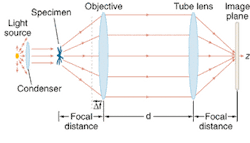
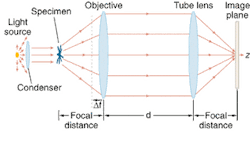
Their optical microscope consisted of a 100× oil-immersion objective with a 1.4 numerical aperture corrected to conjugate the object at infinity, and a tube lens to form the image (see figure). Defocusing consisted of displacing the objective lens. The specimen was illuminated by a tungsten lamp without filters, but the researchers still applied principles of coherent optics because of a very small specimen size on the order of tens of microns. The index of refraction of the specimen was taken as the mean refractive index over the source wavelengths, and they described image contrast mathematically as "proportional to the two-dimensional Laplacian of the phase difference introduced by the phase object and proportional to the amount of defocusing."
They tested the model by baking accumulations of 1-µm-diameter polystyrene beads into spherical caps of varying curvatures and imaging them in both air and water to find good agreement with theory. They then used the defocused microscope to observe an actual biological event, in which they used an optical tweezers (the collimated beam of an infrared laser) to feed a zymosan yeast pellet to a macrophage taken from a mouse. They also repeated the process after effectively sedating the macrophage with the drug cytochalasin D. In both cases, the pellet's demise was filmed through the defocused microscope objective using a digital camera and videocassette recorder. The movies were then digitized using a frame grabber and stored in a computer for analysis.
In addition to noting that it takes about 40 times longer for a drugged macrophage to consume a yeast pellet than it does for a sober macrophage, Mesquita's team also noted two distinctly different dynamic curvature patterns on the macrophage surface. Small, Gaussian-like wave patterns with amplitudes on the order of 0.2 µm seemed to randomly permeate the entire cell surface, while larger, coherent wave patterns with amplitudes on the order of 2 µm appeared to travel from the periphery of the macrophage towards the nucleus. The larger structural undulations increased slightly right around the yeast pellet as it was being swallowed. They also tended to shrink and slow down a good bit after the macrophage had been drugged.
While the rippling behavior of macrophages has been known for some time, the availability of a relatively simple and direct method of dynamic viewing could potentially facilitate the study of cellular processes.
REFERENCE
- U. Agero et al. Physical Rev. E 67, 051904-1 (2003).