Nanosensors enable portable detectors for environmental and medical applications
The development of biosensors for the diagnosis and monitoring of diseases, drug discovery, proteomics, and the environmental detection of pollutants and/or biological agents is an extremely significant problem.1 Fundamentally, a biosensor is derived from the coupling of a ligand-receptor binding reaction to a signal transducer. A variety of biosensor signal transduction methods exist, including optical, radioactive, electrochemical, piezoelectric, magnetic, micromechanical, and mass spectrometric.
One type of optical biosensor detects changes in local refractive index by monitoring the localized surface-plasmon resonance (LSPR) λmax extinction maximum of noble metal nanoparticles with ultraviolet-visible (UV-visible) spectroscopy.2-5 Biosensing with LSPR spectroscopy is exactly analogous to wavelength-shift surface-plasmon-resonance (SPR) spectroscopy.6
Although the absolute areal mass sensitivity of LSPR spectroscopy is almost 100 times less than that of SPR spectroscopy, it has several important characteristics that recommend its study and application to biosensing. The areal mass sensitivity of LSPR spectroscopy is achieved because the short-range distance dependence of the local electromagnetic field, Lz, of 5 to 6 nm, compensates for the modest refractive-index sensitivity of silver (Ag) nanoparticles.
From the instrumentation perspective, LSPR nanosensors can be implemented using extremely simple, small, light, robust, and low-cost equipment for unpolarized, UV-visible extinction spectroscopy in transmission or reflection geometry. LSPR biosensing will be enhanced by and is naturally compatible with handling microfluidic samples. The simplicity of instrumentation afforded by LSPR biosensors should greatly facilitate field-portable environmental or point-of-service medical diagnostic applications.
Nanosensor development
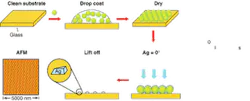
Nanosphere lithography (NSL) is a nanofabrication technique that readily allows the manipulation of the shape, size, and composition of surface-confined nanoparticles (see Fig. 1).7 In our experiments, Ag nanotriangles with in-plane widths of about 100 nm and out-of-plane heights of 50 nm were synthesized using this technique, and their optical properties were monitored using LSPR spectroscopy. The wavelength of the extinction maximum of the nanoparticles shifts as adsorbates induce local refractive-index changes near the surface of the nanoparticles.
FIGURE 2. A tapping-mode AFM image of the LSPR nanobiosensor shows the Ag nanoparticles, top left, (nanosphere diameter, D = 390 nm; mass thickness, dm = 50.0 nm Ag on a mica substrate, scan area, 3.0 µm2); after solvent annealing, the resulting nanoparticles have in-plane widths of about 100 nm and out-of-plane heights of about 51 nm. The surface chemistry of the Ag nanobiosensor (top, right) includes a mixed monolayer of 11-mercaptoundecanoic acid (1) and 1-octanethiol (2). Biotin is covalently linked to the carboxyl groups (3). The anti-biotin binds to a biotinylated Ag nanobiosensor fabricated by NSL on a mica substrate (bottom).
To prepare the LSPR nanosensor for biosensing, the Ag nanotriangles are first functionalized with a self-assembled monolayer. Next, biotin is covalently attached to the carboxylate groups using a zero-length coupling reagent (see Fig. 2).
Sensitivity
The well-studied biotin/streptavidin system, with its extremely high binding affinity (Ka ~1013 M-1), and the antigen-antibody couple, biotin/anti-biotin, were chosen to illustrate the attributes of these LSPR-based nanoscale affinity biosensors (see Fig. 3). The lines are not a fit to the data. Instead, the line was computed from a response model. It was found that this response could be interpreted quantitatively in terms of a model involving 1:1 binding of a ligand to a multivalent receptor with different sites but invariant affinities, and the assumption that only adsorbate-induced local refractive index changes were responsible for the operation of the LSPR nanosensor.
The binding curve provides three important characteristics regarding the system being studied. First, the mass and dimensions of the molecules affect the magnitude of the LSPR shift response. Comparison of the data with theoretical expectations yielded a saturation response, ΔRmax = 26.5 nm for streptavidin (a 60 kDa molecule) and 38.0 nm for anti-biotin (a 150-kDa molecule). Clearly, a larger mass density at the surface of the nanoparticle results in a larger LSPR response. Next, the surface-confined thermodynamic binding constant Ka,surf can be calculated from the binding curve and is estimated to be 1 × 1011 M-1 for streptavidin and 4.5 × 107 M-1 for anti-biotin.
These numbers are directly correlated to the third important characteristic of the system, the limit of detection (LOD). The LOD is less than 1 pM for streptavidin and 100 pM for anti-biotin. As predicted, the LOD of the nanobiosensor studied is lower for systems with higher binding affinities such as for the well-studied biotin-streptavidin couple and higher for systems with lower binding affinities as seen in the anti-biotin system.
Reversibility
For LSPR nanobiosensors to fulfill their mandate, they must be biocompatible and work under physiological conditions. Some binding interactions such as poly-L-lysine to a negatively charged surface can interact reversibly, while other couples with higher surface-binding affinities interact irreversibly. A commercially viable nanobiosensor should be entirely reusable. In the case of this study, this means that the analyte detection must be entirely removable rendering the sensor reusable.
Reusability has a large impact on cost effectiveness and the simplicity of use of biosensors. The reversibility of the binding between anti-biotin and biotin was tested experimentally by exposing an anti-biotin functionalized sample to an excess of concentrated biotin (1 mM in 10-mM buffer). The empty binding sites on the anti-biotin molecules make the molecule susceptible to removal in these conditions (see Fig. 4)
FIGURE 4. An LSPR sensing experiment shows the adsorption/desorption of 20-nM anti-biotin to biotinylated Ag nanoparticles. Anti-biotin is adsorbed onto biotinylated nanoparticles (top): LSPR spectrum of biotinylated Ag nanoparticles in buffer, λmax = 671.1 nm (1); and LSPR spectrum of Ag nanoparticles after incubation in 20 nM anti-biotin in buffer, λmax = 681.8 nm (2). Anti-biotin is desorbed from biotinylated nanoparticles (bottom): LSPR spectrum of biotinylated Ag nanoparticles after incubation in 20-nM anti-biotin in buffer, λmax = 681.8 nm (1); and LSPR spectrum of Ag nanoparticles after incubation in an excess of biotin in buffer for 30 seconds, λmax = 670.2 nm (2).
Considering sensor limitations, the resulting blue-shifted value of -11.6 nm for λmax is essentially identical to its value before treatment with anti-biotin, suggesting that the analyte layer had been completely removed. All measurements were made in 10-mM buffer to ensure the applicability of the results to biological sensing.
Selectivity
Although LSPR spectroscopy is a totally nonselective sensor platform, a high degree of analyte selectivity can be conferred using the specificity of surface-attached ligands and passivation of the sensor surface to nonspecfic binding. For this reason, a set of control experiments was performed to show that streptavidin and anti-biotin binding to the sensor surface containing no capture ligand (biotin), prebiotinylated streptavidin binding to a sensor surface with biotin, and bovine serum albumin in large excess (simulating a clinical sample) binding to a sensor surface with biotin, all produce wavelength shift responses less than that corresponding to the LOD.
Outlook
Extrapolation of our current data indicates that by optimizing these size and shape tunable-nanosensor materials and using single-nanoparticle spectroscopic techniques, it will be possible to reach sensitivities of a few molecules, perhaps even a single molecule, per nanoparticle sensor element. Also, we can reduce the time scale for real-time detection and the study of protein-binding kinetics by two to three orders of magnitude since nanoparticle sensor elements will operate in radial rather than planar diffusion mass-transport regime. In addition, we can implement massively parallel bioassays for high-throughput screening applications while maintaining extremely low sample-volume requirements.
Finally, LSPR nanosensors can be implemented using extremely simple, small, light, robust, low-cost equipment for unpolarized, UV-visible extinction spectroscopy in transmission or reflection geometry. The instrumental simplicity of the LSPR nanosensor approach is expected to greatly facilitate field-portable environmental or point-of-service medical diagnostic applications.
ACKNOWLEDGMENTS
We acknowledge support of the Nanoscale Science and Engineering Initiative of the National Science Foundation under NSL Award EEC-0118025. Any opinions, findings and conclusions or recommendations expressed in this material are those of the author(s) and do not necessarily reflect those of the National Science Foundation. We also acknowledge the support of the Air Force Office of Scientific Research Multidisciplinary University Research Initiative program (F49620-02-1-0381).
REFERENCES
- A. P. F. Turner, Science 290, 1315, (2000).
- A. J. Haes and R. P. Van Duyne, J. Am. Chem. Soc. 124, 10596 (2002).
- A. J. Haes and R. P. Van Duyne, Mat. Res. Soc. Symp. Proc. 723, O3.1.1 (2002).
- M. D. Malinsky, K. L. Kelly, G. C. Schatz and R. P. Van Duyne, J. Am. Chem. Soc. 123, 1471 (2001).
- J. C. Riboh, A. J. Haes, A. D. McFarland, C. Ranjit Yonzon and R. P. Van Duyne, J. Phys. Chem. B 107, 1772 (2003).
- L. S. Jung, C. T. Campbell, T. M. Chinowsky, M. N. Mar and S. S. Yee, Langmuir 14, 5636, (1998).
- C. L. Haynes and R. P. Van Duyne, J. Phys. Chem. B 105, 5599 (2001).
AMANDA HAES is a graduate research assistant in chemistry and RICHARD VAN DUYNE is a professor of chemistry at Northwestern University, Evanston, IL 60208-3113; e-mail: [email protected].