Lasers for Biophotonics: Lasers meet changing demands of biomedical applications
Most laser life science applications fall into one of three categories: cytometry, microscopy, or sequencing, and virtually all of them use fluorescence. As a result, key trends in these areas dominate the development of lasers for the life sciences market.
In this article, we will explore the background, current status, and key trends to paint a picture of the current state of laser technology for bio-optical applications.
True UV in flow cytometry
In flow cytometry, cells are tagged with fluorescent probes and are passed through the interaction zone, where laser beams excite fluorescence. The signal from each cell is separated into several wavelength bands by dichroic beam splitters, prior to detection.
A key trend in research instruments is the increasing use of ultraviolet (UV) lasers to excite endogenous (natural) fluorescence and/or for expanded multicolor analysis/acquisition. For example, endogeneous fluorescence can be used as the sole criterion for sex sorting of sperm; the shorter Y sex chromosome of the male sperm produces slightly less UV-excited fluorescence than the longer X chromosome of the female sperm. Endogenous fluorescence is also attractive in many other sorting applications because it involves no chemical intervention with fluorescent probes. Plus, UV lasers can also expand the total wavelength bandwidth of a flow cytometer in multicolor assays.
The shift from bulky gas lasers to solid state alternatives presented a significant challenge for the ultraviolet. Efficient frequency tripling of the 1064 nm fundamental (to produce 355 nm) requires high intensity at the harmonic-generating crystals, necessitating intracavity tripling in continuous wave (CW) lasers. However, competition for gain among different longitudinal modes leads to chaotic amplitude noise for intracavity doubling and tripling, called "green noise."
This green noise can only be completely eliminated through single-mode operation, but the cost and complexity often cannot be justified for multilaser OEM instrumentation applications. Instead, some manufacturers used extra-cavity tripling of passively mode-locked lasers that have pseudo-CW output with extremely high peak power. Of concern, there are questions about the potential negative impact of using high peak power UV on live cells.
Optically pumped semiconductor laser (OPSL) technology is excellent for generating true-CW ultraviolet because it is does not suffer from green noise. The reason is that the upper state lifetime of the semiconductor gain medium is considerably shorter than the round trip time of the laser cavity, so there is no stored gain to support dynamic mode competition. OPSL technology is also power scalable. As a result, true-CW, OPSL lasers at 355 nm now offer output powers exceeding 20 mW.
Multiple wavelengths in flow cytometry
Another trend in flow cytometry is the simultaneous use of an increased number of fluorochromes to support more complex analyses. For example, ten and more color assays have been shown to provide the sensitivity and specificity for important applications such as determining minimal residual disease (MRD) analysis in various types of leukemia. One approach to increasing the number of fluorochromes that can be independently sensed is to use fluorochromes with narrower emission bands, i.e., less spectral overlap.
Another approach is to increase the instrument's spectral bandwidth, which can then be divided between more individual fluorescence channels. The excitation bandwidth can be expanded by use of UV lasers, or by filling in key gaps, such as the one between 561 and 630 nm. These gaps occurred because of the limited available wavelengths from DPSS lasers and laser diodes.
OPSL technology addresses this because it is wavelength scalable; the fundamental wavelength output by the semiconductor gain chip can be customized by changing the composition and size of the active quantum wells. As a result, OPSLs have been developed at several new strategic wavelengths in the yellow and orange, specifically 552, 577, and 588 nm, as well as 568 nm (which was previously available but only from bulky krypton lasers).
Near-infrared OPSLs (e.g., 833 nm) are also of interest for extending instrument bandwidth on the longer wavelength side. All these wavelengths are available in plug and play formats such as the Coherent OBIS series, simplifying integration and tasks such as hot swapping.
Lasers optimized for super-resolution microscopy
In microscopy, the diffraction limit was long considered unsurmountable, limiting resolution to around 200 nm in the xy plane and 500 nm in the z axis of confocal microscopes. The 2014 Nobel Prize in Chemistry—jointly awarded to Eric Betzig, Stefan W. Hell, and William E. Moerner for pioneering super-resolution microscopy techniques—proves how different and complementary techniques have surpassed the diffraction limit.
All optical super-resolution or nanoscopy techniques involve optically and reversibly preparing states of a fluorescence label that differ in their emission characteristics (e.g., a bright ON and a dark OFF state). Optical nanoscopy methods can be loosely divided into two groups: those that directly improve microscope effective spatial resolution by deterministically photoswitching labels in space and time and those that achieve the higher resolution by stochastically switching single-molecule fluorescence on and off in space.
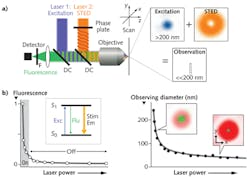
Examples of the first group include stimulated emission depletion (STED) microscopy, ground-state depletion (GSD) microscopy, and reversible saturable optical fluorescence transition (RESOLFT) microscopy. These techniques use two overlapping laser beams. The first beam bleaches fluorescence from a part of the area excited by the second probe beam. The easiest to visualize is STED, which uses a donut-shaped beam. The extent of bleaching, and hence the constriction of the probed area, is varied by changing the power of the STED beam (see Fig. 1).
Examples of stochastic switching methods include (direct) stochastic optical reconstruction microscopy, known as (d)STORM and (fluorescence) photoactivation localization microscopy, known as (f)PALM. Here, extensive sample bleaching by a wide-field light source leaves just a few isolated unbleached molecules.
The samples are imaged by a low-noise camera and algorithms then convert each isolated "blob" into a point location, based on the optical transfer function of the microscope. Repeating the on/off reversible bleaching process results in another random set of un-overlapped blobs. Eventually an entire image is built up of the point locations.
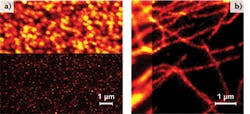
These applications require new wavelengths, particularly orange and yellow, for photobleaching without crosstalk with the fluorescence excitation wavelengths. They also need low noise and higher power—as high as 2 W at the laser output—and the ability to adjust the power to optimize the various super-resolution bleaching mechanisms. OPSL technology is ideal for this purpose; unlike other solid state lasers, OPSLs do not suffer from thermal lensing, so the power can be dynamically adjusted (from 10% to 100% power) with no effect on beam quality and pointing. Figure 2 shows gated STED images acquired with one of these yellow OPSLs at 577 nm.Divergent multiphoton microscopy techniques
Multiphoton microscopy is another dynamic area because of the continued development of diverse methods. Key needs here are femtosecond lasers that combine versatility and industrial reliability for multiuser facilities, higher power at wavelengths longer than 1 μm, and dual-wavelength capability to support exciting developments in optogenetics, in which light-sensitive proteins are used to optically stimulate/inhibit specific neurons. A second laser wavelength is then used to map activity in connected neurons using two-photon excitation of calcium indicators.
While Ti:sapphire lasers continue to be the workhorse sources for multiphoton microscopy, femtosecond lasers based on ytterbium technology have recently gained popularity. Used directly at their wavelength of 1055 nm, they excel in activating red-shifted fluorescent proteins, photoactivators, and Ca indicators. They can also be used to pump an integrated OPO to achieve broader tunability (680–1300 nm) than Ti:sapphire lasers.
The combination of a 1–2 W, 1055 nm fixed output with the tunable output of the OPO or a Ti:sapphire laser perfectly addresses optogenetics, where dual-color excitation can independently excite/inhibit and detect action potentials in neurons. These dual-wavelength sources are also well-suited to other methods, including coherent Raman techniques like coherent anti-Stokes Raman scattering (CARS) and stimulated Raman scattering (SRS).
Lasers for DNA sequencing and profiling
Automated sequencing and profiling are still in their infancy, and sequencing methods vary significantly among instrument developers. Most of these utilize laser-excited fluorescence, but the challenge for our industry is that laser performance (beam quality, wavelength, and power) packaging, and cost requirements are as widely different as the techniques themselves.
A comprehensive survey of sequencing methods is beyond the scope of this article. (For a more detailed discussion, see "Laser fluorescence powers sequencing advances" in the January/February 2015 issue of BioOptics World.) Here, we present a brief overview designed to show some of the technical diversity.
The successful Human Genome Project relied on automating the so-called Sanger method, in which DNA is first cut into strands that are a few hundred bases long. These are then copied in vitro using DNA polymerase in the presence of normal bases (adenosine, cytosine, guanidine, and thymidine) as well as a small amount of chemically modified bases that "jam" the synthesis process when incorporated into the new copy. Each of these modified bases is also labeled with a different fluorophore for each base type (ACGT).
The result is a mix of every possible length, from a few bases all the way to the original hundreds of bases, with a fluorescent label identifying what base it ends with. These are then run through a capillary electrophoresis column to separate them according to length; the column exit is excited with a focused 488 nm laser. The sequence is then read by recording which fluorophore appears next and so on. Only a few hundred bases can be read in this way, so sequencing the entire human genome (~109 bases, circa 35,000 genes) was a massive collaborative challenge costing roughly $3 billion.
NGS and emerging platforms
In just 20 years, the cost per entire human genome has experienced a reduction of seven orders of magnitude. So-called next-generation sequencing (NGS) and emerging third-generation instruments have made this possible by the ability to simultaneously sequence up to hundreds of thousands of lengths of DNA and/or to analyze longer cut strands.
The most popular NGS attaches myriad cut strands-to be analyzed on a lab-on-a-chip setup-to a series of micron-sized spheres that are held in a 2D array by a pattern of tiny wells on a glass surface. These are then amplified (cloned) so that each strand becomes a tightly localized cluster of identical strands. They are then sequentially exposed to a DNA synthesizing agent (polymerase) together with alternately applied fluorescently labeled bases and removable terminating agents. All four base types are identically labeled.
Initial mapping with a 660 nm laser is followed by excitation with a 532 nm laser together with imaging using a low-noise camera. Each dot in the image flashes when it picks up a fluorescent base, enabling a computer to record the sequence. Software then analyzes all the random strands, with read lengths up to thousands of bases, using sequence overlaps to assemble the sequence of the original uncut length of DNA.
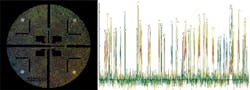
One particularly interesting method with novel optics has been developed by Pacific Biosciences. This system uses zero-mode waveguides (ZMWs), optical waveguides whose diameter is much smaller than the wavelength of light. On a so-called single-molecule real time (SMRT) chip, ZMWs are created by patterning circular holes (~ 70 nm diameter) in an aluminum film (100 nm thickness) deposited on a clear silica substrate. A single polymerase enzyme is located at the bottom of each waveguide (see Fig. 3).Laser illumination occurs through the silica and can only penetrate about 30 nm into each waveguide, so millisecond pulses of fluorescence can only be efficiently collected from the nucleotide as they are being added to the DNA. Importantly, in the company's PacBio RS II system, up to 150,000 sequencing reactions are monitored simultaneously, in real time, by dividing the lasers into that many individual diffraction-limited spots targeted to the individual ZMWs. Moreover, the company claims single continuous read lengths as long 40,000 bases.
A wide number of methods are also targeted at profiling. Rather than recording every individual base, these methods look for specific sequences, for applications ranging from epidemiology to forensics and pre-natal screening. One clever method here first inserts fluorescently labeled target sequences, which are incorporated within the DNA double helix. The treated samples are forced through ultra-narrow channels on microarray plates so that the DNA tertiary structure is stretched to leave a string-shaped molecule with various different fluorescent points along its length. These can be read like a colored barcode using a laser and detector.
In terms of laser power, requirements vary from a few milliwatts for techniques using a tight focus to multiple watts for techniques using wide-field illumination. Similarly, some applications need a perfect TEM00 output whereas others work with a lower cost, multimode laser. And, several of these highly automated techniques have run times of several hours or even days, so long-term power and pointing stability are very important. Reliability is also critical, as a laser problem will ruin the entire data run.
Preferred laser wavelengths are still dominated by argon ion and DPSS legacy wavelengths (488, 514, 532 nm). But the wavelength scalability of OPSL technology is creating interest in new wavelengths, such as 640 nm. In particular, the development of 505 nm OPSLs is potentially disruptive; as the arithmetic mean of 488 and 514 nm, this new wavelength can be used to simultaneously excite some of the fluorophores previously optimized for these two separate wavelengths, simplifying instrument design and lowering instrument costs.
About the Author
Marco Arrigoni
Marco Arrigoni is vice president of marketing at Light Conversion (Vilnius, Lithuania). He previously served as director of marketing at Coherent (Santa Clara, CA) from 2007 through 2023.
Nigel Gallaher
Product Manager, Coherent
Nigel Gallaher is a Product Manager at Coherent (Santa Clara, CA).
Dan Callen
Product Line Manager, Direct-diode Laser Systems at Coherent
Dan Callen is Product Line Manager for Direct-diode Laser Systems at Coherent (Santa Clara, CA).
Darryl McCoy
Director of Product Marketing and Senior Product Line Manager, Coherent
Darryl McCoy is Director of Product Marketing and Senior Product Line Manager at Coherent (Santa Clara, CA).
Volker Pfeufer
Senior Product Line Manager, Coherent GmbH
Volker Pfeufer is Senior Product Line Manager for Diode-pumped Solid State Lasers at Coherent GmbH (Luebeck, Germany).
Matthias Schulze
Director of Marketing, OEM Components and Instrumentation at Coherent
Matthias Schulze is Director of Marketing, OEM Components and Instrumentation at Coherent (Santa Clara, CA).

