Live Cell Imaging/Fluorescence Microscopy: On-the-fly optical superresolution in one simple step
THOMAS HUSER, MARCEL MÜLLER, and ANDREAS MARKWIRTH
Superresolution optical microscopy has come of age and is now available in a number of commercial implementations. For the most part, however, those who use these systems pay a price for achieving such fine optical resolution: The imaging process typically requires considerably more time than does conventional microscopic imaging. This shortcoming was recently addressed by researchers at the University of Bielefeld (Bielefeld, Germany), as well as the Leibniz Institute for Photonic Technologies and Friedrich Schiller University (both in Jena, Germany). The team has developed a novel imaging system that generates superresolution imagery instantaneously, on the fly.1
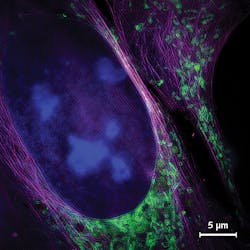
To achieve this, the research team used superresolution structured illumination microscopy (SR-SIM), which, in its 2D implementation (there is a 3D implementation as well), requires acquisition of just nine source images to produce a single final image with approximately 130 nm spatial resolution (see Fig. 1). The researchers call their SR-SIM approach VIGOR (Video-rate Immediate GPU-accelerated Open-Source Reconstruction).
Projecting patterns
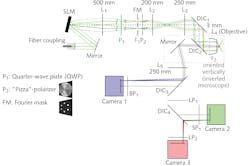
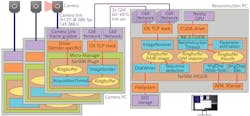
For each of the nine source images, the researchers use an interference pattern with a periodicity at the very edge of what an optical microscope can resolve. This pattern is projected onto the specimen to excite fluorescence in agents they previously applied. To achieve space-filling illumination, the system projects three images with different phases—and at three different angles, each offset by 60 degrees—using a spatial light modulator (SLM) as a switchable, binary optical grating to generate the pattern. The patterns are illuminated by a 12-mm-diameter collimated laser beam from an argon-krypton (ArKr) laser (see Fig. 2).The entire SR-SIM setup can be controlled through a single graphical user interface that enables simple, intuitive operation. The platform provides a user experience similar to that of a standard wide-field fluorescence setup, thanks to the design, which integrates time-critical image acquisition with automatic, real-time image reconstruction and display. Users can select areas of interest, focus in on them, acquire superresolution multicolor imagery at speeds faster than video rate, and share the experience with other interested parties as the process is happening.
Biological applications
To demonstrate on-the-fly reconstruction of images depicting biological samples, the team turned to fixed human bone osteosarcoma epithelial cells (from the U2OS line). They used fluorescence labeling to stain especially fine structures such as mitochondrial membranes and the endoplasmic reticulum that are not resolvable with wide-field microscopy. They showed how the wide-field view responds immediately when the specimen shifts or focus is adjusted, and how the image display is generated with a delay of 250 ms or less. They also used the superresolved images to check particular regions of interest within the cells.
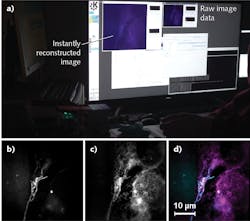
To validate the system’s ability to perform live-cell imaging, the researchers used MitoTracker green to stain mitochondria in living U2OS cells. A broad range of scientists see the ultrastructures of the organelles—again, not resolvable using conventional wide-field microscopy—as having potential to reveal relationships between mitochondrial function and structural changes. The team was able to image organelle dynamics at sub-second frame rates using individual exposure times of just 5 ms/frame, enabling them to avoid motion blur over prolonged timeframes. Figure 4 shows excerpts from such a time-lapse acquisition and highlights the dynamics. The mitochondrial membrane, including folds within the inner membrane called cristae, are indistinguishable in the wide-field images, but clearly represented in the SIM images.The researchers were able to observe SIM reconstructions, including those depicting mitochondrial dynamics, in real time as the experiments were underway. This enabled them to interact directly with the experiment—for instance, to transition among regions of interest, record particular events, and even to discard results—all without waiting for post-processing. These capabilites completely transform the SR-SIM imaging workflow, condensing two separate steps into one. The original two-step process involved acquisition of raw data (with limited feedback during the acquisition phase), followed by reconstruction of images on a dedicated workstation. In the single-step integrated approach, full super-resolution data are available immediately to those running the experiment.
Flexible and freely available
The new, integrated system provides great flexibility, allowing scientists to quickly adjust imaging conditions and, for instance, select illumination duration and between-frame delay for time-lapse image capture. Such options are especially useful for limiting photodamage in live-cell experiments. For instance, a user could browse a specimen in a low-damage mode (capturing perhaps one image every second with fast exposure to alleviate motion blur), and then, upon finding a specific area with dynamics of interest, switch to using full frame-rate and high speed. Such processes might even be automated, and, with help from the researchers' instant image processing, enable a type of intelligent microscopy.
The researchers have made the details of their work freely available to the community under the fairSIM project (fairsim.org), which provides free and open-source tools and resources, and under open-source license. They have shared not only their optomechanical design for SR-SIM, but also multicolor extensions, plans for electronic and software control components, and reconstruction software. Through this means, they hope to encourage others to copy and adapt their methods—from upgrading existing SR-SIM microscopy systems with on-the-fly reconstruction to developing other SR-SIM implementations—for instance, nonlinear SIM, total internal reflection fluorescence (TIRF) SIM with the highest NA objectives available, and perhaps even detection schemes involving multiple planes.
REFERENCE
1. A. Markwirth et al., Nat. Commun., 10, 4315 (2019); doi:10.1038/s41467-019-12165-x.
Thomas Huser, Ph.D., is Professor of Physics and leads the Biomolecular Photonics Research Group, Marcel Müller, Ph.D., is a Postdoctoral Associate, and Andreas Markwirth is a PhD candidate, all at the University of Bielefeld, Bielefeld, Germany; e-mail: [email protected]; www2.physik.uni-bielefeld.de/542.html?&L=1.