Novel microscope for dynamic structural biology with single-molecule FRET
The confocal fluorescence microscope has become a workhorse instrument routinely used in biology to visualize dynamic processes and cellular structures. Beyond imaging, a confocal microscope can provide point measurement techniques such as fluorescence correlation spectroscopy (FCS) or single-molecule fluorescence resonance energy transfer (smFRET), which are gaining popularity for biophysical studies.
One more advance that has received attention recently is the addition of fluorescence lifetime imaging (FLIM). Measuring the fluorescence lifetime of a fluorophore opens an extra dimension of information, which is complementary to intensity and spectral information. The fluorescence lifetime is a characteristic of each fluorophore and its environment. It can be influenced, for example, by the smFRET process, such that the shortening of the donor lifetime in the presence of an acceptor reports on the FRET efficiency.
PicoQuant’s new microscope, Luminosa, is designed for best performance within the lifetime domain. Moreover, it’s simultaneously optimized for single-photon sensitivity, enabling experiments at the single-molecule level.
The variety of time-resolved methods offered by the microscope are suitable for a broad range of applications in life sciences, such as dynamic structural biology, study of cellular mechanisms driven by phase separation, or mapping dynamics and structure of cellular membranes.
Across many research areas, scientists regularly have trouble reproducing other scientists’ experiments.1 One potential way to increase accuracy, precision, reproducibility, and quality of experiments is to limit human error by automation of microscopy. Additionally, this increases the ease of use of imaging systems, all in all saving researchers time and materials. Moreover, automation boosts the throughput, enabling new experimental possibilities such as time-lapse imaging, multiposition imaging, or multipoint time traces.
Intuitive workflows for easy day-to-day operation
Luminosa’s software permits quick and easy workflows for experiments, allowing users to focus on their samples. It incorporates GPU-accelerated algorithms for smFRET, FCS, and time-resolved imaging methods. Its high speed enables automated analysis routines, which takes information about the experiment type, hardware configuration, fluorophores, and data dimensions into account. So, initial results are obtained without user interaction. The software also offers the possibility to flexibly define custom measurement and analysis modes.
Based on single-photon counting, Luminosa’s optical design is optimized for highest sensitivity by incorporating only the minimal number of optical elements in the best quality. It includes the latest generation of PMA Hybrid or SPAD detectors and time-correlated single-photon counting (TCSPC) electronics for time-resolved measurements.
Use case: smFRET for dynamic structural biology
Single-molecule studies and more specifically smFRET methodologies have become a standard tool for studying dynamic structural changes in proteins and nucleic acids. They can reveal dynamic events on timescales from nanoseconds to seconds. This allows observing, for example, chain dynamics, binding, folding, allosteric signaling, and oligomerization. The results nicely complement structural data obtained with other methods such as electron microscopy or nuclear magnetic resonance (NMR), so they can now be entered into the archiving system for integrative structural models, PDB-Dev.2 The power of smFRET is highlighted by the study of intrinsically disordered proteins, where other methods for structure determination reach their limit.
smFRET measurements can be performed with molecules freely moving in solution as well as with molecules immobilized on the coverslip surface. For each modality, Luminosa offers a dedicated workflow. To start out, the researcher chooses which measurement technique to use in the next experiment. The system can quickly perform a sample-free auto-alignment, ensuring that it is always in the same good condition. A clearly arranged software interface shows only parameters necessary for the type of measurement, minimizing the risk of accidental errors.
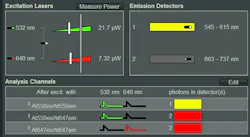
Upon selection of the FRET pair in use, the hardware is automatically configured correctly—that is, excitation wavelengths and power are set, pulsed interleaved excitation of both donor and acceptor is enabled, and correct detectors and emission filters are chosen (see Fig. 1). Moreover, the analysis channels are assigned based on the experimental context: donor emission after donor excitation, sensitized acceptor emission after donor excitation, and acceptor emission after direct acceptor excitation as control.
Similar functionality is already available in most commercial laser scanning microscopes, but here additional parameters important for time-resolved measurements, such as laser repetition rate, are now set as well, reducing the amount of expert knowledge required from the instrument user. Saving and loading of settings guarantees the repeatability of measurements. Furthermore, the appropriate metadata is saved alongside the actual data, and can even be searched later. The fast excitation laser power calibration allows setting and displaying the excitation intensity in microwatts (see Fig. 1), without requiring an external power meter.3 This underlines the importance of illumination power in reproducibility, as photobleaching, phototoxicity, etc. cause variability of results.
Different stop conditions for the acquisition can be selected, depending on the measurement type. For example, for molecules in solution, these are the number of bursts, total time, or reduction of burst rate. This promotes collection of consistent datasets with adequate statistics.
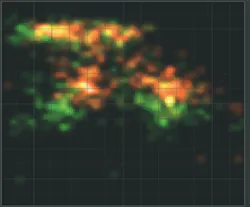
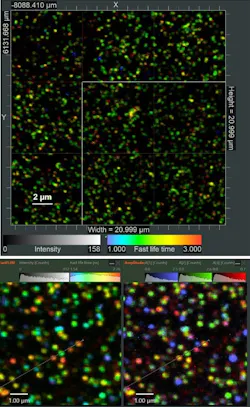
While the measurement is running, several online previews are displayed in real time. In the case of smFRET in solution, these are a histogram of FRET efficiency (E) vs. stoichiometry (S), a burst histogram, an intensity time trace, and a TCSPC histogram with online fitting of the donor and acceptor lifetime decays. The values of E and S are corrected with four factors, according to the standard procedure of the community.4 Both corrected and uncorrected data are shown so that the influence of the correction procedure can be assessed (see Fig. 2). The preview allows researchers to immediately judge the quality of the sample, and to ascertain the correct settings of acquisition parameters. Users can manually change the correction factors later, because the raw data is always saved unchanged.After acquisition of an image tile, an algorithm automatically detects the positions of single molecules according to selectable criteria. Then, the molecules are targeted one after the other and intensity time traces are recorded from each until the pre-defined stop criteria are met. Afterward, the next image tile is scanned. In this way, datasets with adequate statistics can easily be obtained.
Luminosa enables researchers to easily apply single-molecule and time-resolved fluorescence microscopy methods in a wide range of life and materials science studies, without requiring years of microscopy expertise. Thanks to many new software features in combination with state-of-the-art hardware, the system quickly delivers high-quality data in a reproducible manner. Specifically for dynamic structural biology studies, Luminosa allows many researchers to view their samples from a new angle.
REFERENCES
1. M. Baker, Nature, 533, 452–454 (2016).
2. E. Lerner et al., eLife, 10, e60416 (Mar. 29, 2021); doi:10.7554/elife.60416.
3. P. Montero Llopis et al., Nat. Methods, 18, 1463–1476 (2021).
4. B. Hellenkamp et al., Nat. Methods, 15, 669–676 (2018).
5. G. Agam et al., bioRxiv (2022); https://doi.org/10.1101/2022.08.03.502619.
6. T. Niehörster et al., Nat. Methods, 13, 257–262 (2016).
7. S. Rohilla et al., Sci. Rep., 10, 3820 (2020).
About the Author
Maria Loidolt-Krüger
Scientific Content Creator, PicoQuant
Maria Loidolt-Krüger is scientific content creator at PicoQuant (Berlin, Germany).