NANOMETROLOGY: QI offers next-generation AFM imaging mode for nanometrology
HEIKO HASCHKE and TORSTEN JÄHNKE
There are many challenges in applying atomic-force microscopy (AFM) to the high-resolution requirements of the modern user. No longer does the original AFM contact imaging mode developed 25 years ago provide the ability to produce images of samples in liquids without damaging the sample. While a variety of so-called noncontact methods have been introduced, they still cannot meet the requirements of researchers in the life sciences where samples are soft and delicate, requiring care in preparation and imaging in a quantitative manner. This user need has directed the developments of JPK Instruments and has led to a new quantitative imaging (QI) mode.
From the beginning of QI development, the design goals were clear: In order for life scientists more familiar with light microscopy operation to accept AFM and its new imaging modes, it was necessary to design a system with quick and intuitive processes that could be used in various environments such as air or liquid. The mode needed to handle "difficult" samples that are soft and/or sticky, perhaps loosely attached to the mounting substrate, or that have particularly sharp features/edges. This meant that the new nanometrology mode should provide extremely precise control of the force of the probe as it interacts with the sample at every imaging pixel.
New imaging modes must also provide quantitative data, and the QI mode offers mechanical, chemical, and electrical data. In addition, the quantitative provision of real-force curves while imaging delivers the maximum amount of data points. The area of cell adhesion, or the measurement of forces between a single molecule and another (or a surface), are rapidly growing application needs when using AFM in the life sciences.
Defining QI
The new QI, which is compatible with standard cantilevers, is a force-curve-based imaging mode where the user has full control over the tip-sample force at every pixel of the image, meaning there is no need for set-point or gain adjustment while scanning. Its tip movement algorithm—TipSaver—prevents lateral forces and controls vertical forces, making nondestructive measurements for cantilever tip and sample possible.
Coupled with ForceWatch technology, imaging problems with soft (hydrogels or biomolecules), sticky (polymers or bacteria), loosely attached samples (nanotubes or virus particles in fluid) or samples with steep edges (powders, MEMS structures) are removed. Essentially, ForceWatch uses a set of algorithms that closely monitor the feedback signal and make very rapid adjustments as signal changes are observed.
The QI mode is particularly useful in areas that demand both high resolution and force sensitivity such as in biology, and in polymer and surface science where samples have challenging physical parameters.
The only parameters that need to be set prior to imaging are the imaging force and the z-length for the acquired force curve per imaging pixel. Both parameters are intuitive and do not require detailed technical knowledge. In addition, both are measured precisely by ForceWatch and our capacitive sensor technology. Complex determination of driving frequencies and voltages with sweeps, amplitude setting, and phase corrections before imaging are obsolete.
There is no feedback loop in QI; therefore, no parameters will have to be set or optimized during imaging. Additional feedback loops such as phaselocked loop (PLL) or amplitude gain control (AGC) are unnecessary. This lack of feedback loops leads to absolutely independent imaging pixels, eliminating streaks caused by just one improper imaging point.
Piezo motion normal to the sample plane (z-axis) has been carefully optimized in QI in order to achieve comparable imaging speeds to all other AFM imaging modes. This optimization in the z direction was required to perform the force curves with reliable approach and retract rates essential for quantitative analysis, such as those required for indentation or adhesion applications.
Most important for the life sciences user, QI can be combined with all common optical imaging techniques such as high-numerical-aperture confocal laser scanning microscopy (CLSM) by using JPK's DirectOverlay approach that combines AFM and optical microscopy. It utilizes special optical calibration and import routines, and displays AFM simultaneously with optical microscopy information from the sample. Interesting locations on the sample can be identified with a wide variety of contrast methods, such as optical phase contrast, differential interference contrast (DIC), or variable relief contrast (VAREL).
In addition, the use of fluorescence techniques such as epi-fluorescence, confocal, or total internal reflection fluorescence (TIRF) microscopy provides insight into the behavior or location of particular molecules or species. Combining AFM imaging or force measurements with these optical methods on the same spot at the same time is not straightforward. The optical image acquisition is independent from the AFM image recording, meaning a tip-scanning AFM design is critical to this application and underscored through the optimal use of closed-loop imaging in the lateral direction (x, y) using capacitive sensors.
Quantifying difficult samples
During QI imaging there are no lateral forces applied to the sample, so imaging of loosely attached objects or even nonimmobilized samples (often found in life science applications) is now possible. The force-based nature of QI allows imaging of taller objects with more sticky surfaces compared to conventional AFM imaging modes. Side-to-side rastering of a probe can sometimes damage or move the surface of such samples. This contrasts with a force-curve measurement that moves the probe up and down vertically, without dragging through the surface.
The QI multiscan feature allows unattended scanning of even centimeter-sized surface areas. This enables much larger areas to be scanned (hundreds of microns), improving the ability to locate, zoom in, and study specific "small" features on the nanometer scale since full data sets of force curves are stored for post-experiment processing by the user.
To give users a choice of capabilities, there is also a QI advanced software package that enables high-spatial-resolution quantitative measurement of nanoscale material properties such as stiffness, adhesion, dissipation, and more. It provides more data channels and features for data extraction and processing to analyze additional sample information. Depending on application, it is also possible to perform electrical conductivity or molecular recognition measurements from a single image.
Advanced controller systems now allow any signal measured in an experiment to be recorded and stored in parallel. Improvements in electronic components have enabled suppliers to offer a digital controller with extremely low noise levels; for example, JPK's Vortis SPM Control Station uses high-speed lock-in amplifier technology to provide precise amplitude and phase detection to enable sharper, more accurate, quantitative imaging.
And finally, all recorded data channels can be plotted as 3D images and—due to the force-spectroscopy nature of QI—as 3D images at different positions normal to the sample surface. For instance, a topography map of zero-interaction force can be plotted to yield more details on the unperturbed sample surface. For example, in JPK's data processing (DP) software, all data channels with full bandwidth at every imaging point are available for more flexibility. The DP software also offers several analysis algorithms for QI data, such as extracting adhesion values or providing Young's modulus information.
Applications
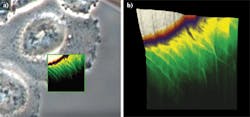
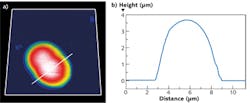
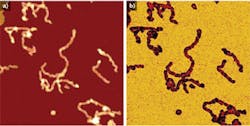
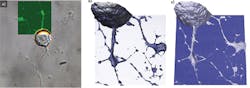
Using a JPK NanoWizard 3 AFM, living Chinese hamster ovary (CHO) cells in HEPES buffer (an organic chemical buffering agent) at zero imaging force were imaged for the first time (see Fig. 1). These images give new insights into the exact cell topography because the independence of every data point removes any scanning artifacts. This QI method was also used to image living and nonimmobilized cyanobacteria, which have also been acquired with zero imaging force in buffer solution (see Fig. 2). The absence of forces gives the precise topography without any image distortion.Looking ahead, it is expected that QI will become a vital tool for the life-science microscopist in making rapid, quantitative, and accurate nanometrology measurements.
Heiko Haschke is head of applications and Torsten Jähnke is founder and chief technology officer of JPK Instruments AG, Bouchéstrasse 12, 12435 Berlin, Germany; e-mail: [email protected]; www.jpk.com.