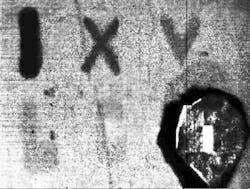PHOTONICS APPLIED: FORENSICS: When photonics meets forensics, crime really doesn't pay
The phrase “crime doesn’t pay” is truer than ever before: Scientific techniques—many of them photonic or optical in nature—have reached a level of sophistication that allows the forensic scientist to definitively link a suspect to a crime scene. There is no “getting away with murder” when the tiniest trace of bodily fluids, hair and clothing particulates, or even soil or vegetation clues are left behind during a traumatic event.
We all know that chemical DNA analysis has been instrumental in solving numerous current and decades-old cold cases. But were you aware of the ability of microspectrophotometry to identify microscopic textile samples, or that laser ablation of a single hair sample (http://opfocus.org/index.php?topic=story&v=8&s=6) can reveal the isotopic ratios of such chemicals as oxygen, nitrogen, sulfur, and carbon present in the keratin to reveal where an individual was living and what they were eating as a function of time?
Television shows such as CSI: Crime Scene Investigation, Bones, and Forensic Files have popularized the science of forensics. Once limited to archaic and destructive chemical and laboratory intensive procedures, the processing of crime scene evidence is now possible using light- and laser-based optical and photonic methods—even when only very minute or microscopic trace evidence is left behind. For example, Raman spectroscopy, which is approximately ten times more sensitive than mid-infrared (IR) spectroscopy, can definitively analyze diluted bodily fluids to a high degree of accuracy without destroying the sample or requiring hazardous chemicals.1
The scene of the crime
In any forensic investigation, the first step is to analyze the crime scene, looking for traces of fluids or solids that could provide clues. While an ordinary “black light” or ultraviolet (UV) light (coupled with filters and goggles to improve visualization) can find naturally fluorescing semen, vaginal fluids, urine, sweat, and saliva, blood stains are a different matter: Blood absorbs all UV wavelengths and can be viewed as a dark stain against a brighter background. Unfortunately, criminals often attempt to hide any trace of bodily fluids through cleaning, rendering UV analysis useless. But high-tech photonics offers a solution: IR spectroscopy.
Even for a bloodstain that has been diluted 100 times through cleaning, IR spectroscopy can be an effective detection tool. Professors Stephen L. Morgan and Michael L. Myrick at the University of South Carolina (Columbia, SC) concluded a National Institute of Justice (NIJ) grant (https://www.ncjrs.gov/pdffiles1/nij/grants/235286.pdf) in June 2011 by developing a technique for “Rapid Visualization of Biological Fluids at Crime Scenes using Optical Spectroscopy.” They point out that for very small volumes of blood, chemical analysis using luminol or other enhancement chemicals such as amido black, fluorescein, or leuco-crystal violet often creates false positives. The reason for this is because fluorescence of these materials is catalyzed not only by the iron in blood hemoglobin but also by any naturally occurring iron at the crime scene as well as other common household materials.
Alternatively, Fourier transform IR (FTIR) spectroscopy can nondestructively and in a noncontact fashion detect the strong absorption of hemoglobin at 1650 and 1540 cm-1 against the nonabsorbing background of common surfaces and textiles.
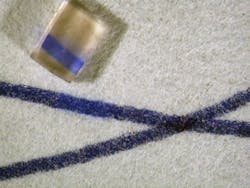
Morgan and Myrick’s instrument uses an IR source (a glow-bar or space heater) combined with a conventional thermal IR camera. The source is chopped and each pixel in the image is digitally processed by a lock-in amplifier to reveal the visual contrast between stain/no-stain regions. The detector response is optimized by a combinatorial, simulation-driven design process to select chemical filters that maximize the discrimination between blood and unstained surfaces. In addition to ready detection of blood using this thermal IR imaging technique, the team is also working to be able to distinguish blood from other interfering compounds by viewing samples through one or more chemical filters composed of polymer films on IR-transparent substrates (see Fig. 1).
Beyond IR imaging and spectroscopy, Raman spectroscopy can provide improved sensitivity in bloodstain and semen detection, but it is also critical for the identification of tiny amounts of unknown substances at the crime scene.
“Raman spectroscopy has several advantages over IR spectroscopy, but the most important is that it can be used in aqueous solvents due to the low Raman scattering of water,” says John R. Lombardi, chemistry professor at the City College of New York (CCNY; New York, NY). As we learned in certain handheld spectrometer applications and as Lombardi reiterates, “Water is a strong IR absorber and interferes with the IR signal, making Raman best for biochemical studies.”
But Lombardi says that there are two distinct disadvantages of Raman spectroscopy: weakness of the Raman signal and interference from fluorescence. Fortunately, both of these weaknesses are overcome by surface-enhanced Raman spectroscopy (SERS), in which a molecule is adsorbed on a metal nanoparticle surface (usually silver or gold). “Using SERS, the Raman signal is enhanced by many orders of magnitude while simultaneously quenching fluorescence,” says Lombardi. “Enhancement factors of up to 1014 have been reported, making possible the detection of a single molecule.2 This makes it ideal for detection and identification of trace quantities of materials, and is the basis for the increasing use of SERS in forensic applications.”
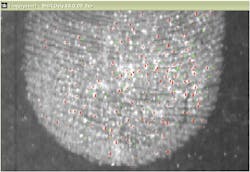
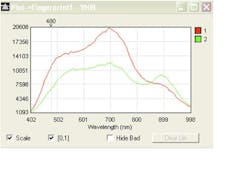
In conjunction with the Metropolitan Museum of Art, the NYPD crime laboratory, and the forensic science program at John Jay College of Criminal Justice (all in New York, NY), Lombardi says that CCNY has been developing techniques to apply SERS to examine trace quantities of substances of interest in forensic science, such as controlled substances (drugs), dyes used in tattoo inks, and inks and dyes used in artwork and textiles.3-6 For example, the team has developed a technique similar to solid phase microextraction (SPME), in which a methacrylate hydrogel and a 1:1 dimethylformamide (DMF) and water solution with 1% weight by volume disodium ethylenediaminetetraacetic acid (EDTA) is used to extract trace amounts of a target colorant from a substrate (see Fig. 2). Once the trace amount of dye is extracted by the hydrogel, it is dissolved in a photonic solution of metal nanoparticles for SERS detection.In cases of controlled substance analysis as part of an autopsy—or alternatively, the analysis of what can turn out to be counterfeit drugs at a crime scene—Raman spectroscopy and its variations are also playing a major role. Senior scientist Pavel Matousek and his colleagues at the Central Laser Facility at STFC’s Rutherford Appleton Laboratory (Oxfordshire, England) developed the spatially offset Raman spectroscopy (SORS) technique that analyzes the Raman signal returned from regions up to 10 mm beyond the initial entry point of a beam into an even opaque sample such as a pill bottle or biological tissue. The instrument uses conventional CW laser excitation and dispersive Raman-CCD detection. The method relies on the fact that Raman signals emerging from deeper areas of turbid samples are more broadly spread on the surface of the sample than those originating from shallower depths—a direct consequence of photon diffusion within the sample.7
Since its development in 2005, SORS has been commercialized through STFC spinout Cobalt Light Systems (Oxfordshire, England). Recent developments include a SORS device for the noninvasive screening of liquid explosives concealed in plastic and glass bottles for aviation security, which has exceptionally low false alarm rates compared with alternative technologies and is currently being tested at major European airports (www.cobaltlight.com/products/insight100). Another commercial instrument under development is a SORS scanner for the noninvasive identification of incoming raw materials in pharmaceutical manufacturing that can establish the identity of chemicals within seconds—without opening the packaging. The SORS technique is also being developed worldwide in a number of academic research laboratories for noninvasive screening of bone diseases and for breast cancer diagnosis.8
DNA fragments, partial fingerprints
Just as the crime scene often reveals only diluted bloodstains and minute traces of other substances, DNA material and fingerprints are rarely complete. However, DNA fragments and partial fingerprints are sometimes enough to identify an individual, especially considering how photonics technology continues to evolve and gain in detection sensitivity.
In 2008, after earlier false identifications based on dental records and preliminary DNA analysis, the identity of a young boy recovered from the Titanic disaster of 1912 and buried in Halifax, Nova Scotia, was finally revealed: His name was Sidney Leslie Goodwin. “The boy’s positive identification was made possible through several iterations of a capillary electrophoresis method of detecting fluorescently labeled DNA markers called single nucleotide polymorphisms [SNPs],” says forensic genealogist Colleen Fitzpatrick from Identifinders International (Huntington Beach, CA), who worked with the Armed Forces DNA Identification Laboratory (Rockville, MD) as part of the team that scientifically identified the remains.9
Detailed DNA analysis uses a variety of photonic methods; principally, the techniques involve fluorescence. Commercially available instrumentation such as the PCR Amplification Kit and SNaPshot from Applied Biosystems (Foster City, CA) use one or more laser sources to excite particular fluorescent dyes or fluorophores that have been attached to DNA strands. Methods that amplify and then measure the fluorescent signals, as well as other photonic sensor techniques, can provide definitive DNA matching using even very small amounts or fragments of DNA material.
“Hyperspectral sensors can identify material and compounds left in the ridges of a fingerprint based on the inherent chemical composition of the sample,” says David Bannon, CEO of Headwall Photonics (Fitchburg, MA).REFERENCES
1. V. Sikirzhytski et al., BioOptics World, 5, 1, 33–36 (January/February 2012).
2. M. V. Cañamares et al., J. Phys. Chem. C, 112, 20295–20300 (2008).
3. M. Leona et al., Analyt. Chem., 83, 3990–3993 (2011).
4. I. Geiman et al., J. Forensic Sci., 54, 947–952 (2009).
5. M. Leona et al., Acc. Chem. Research, 43, 782 (2010).
6. V. Rana et al., J. Forensic Sci., 56, 200–207 (2011).
7. R. Ehrenberg, ScienceNews, 179, 13, 22 (June 18, 2011).
8. K. Buckley and P. Matousek, Analyst, 15, 136, 3039–3050 (2011).
9. R. S. Just et al., Forensic Sci. Int.: Genetics, 5, 3, 231–235 (June 2011).
Building a better database
Forensic data can only be used to catch a criminal if the DNA, blood type, or fingerprint details can be matched to stored information in a database or directly to a suspect. Incredibly, most DNA and fingerprint databases are on the national (and not international or global) level; in fact, the INTERPOL “global” DNA and fingerprint database reportedly contains only 117,000 DNA samples and only 151,000 fingerprint records—far from comprehensive.
In late 2011, the National Institute of Standards and Technology (NIST; www.nist.gov) published a revised biometric standard to extend both the type and amount of information that can be shared by forensic scientists. Called the Data Format for the Interchange of Fingerprint, Facial & Other Biometric Information (see www.nist.gov/customcf/get_pdf.cfm?pub_id=910136), the standard includes DNA, fingerprint, palmprint, footprint, facial recognition, and iris information. But the standard is much more than a collection of images and data; it also defines how to specify and share the geo-positioning coordinates of biometric sample collection to make sure that data analysts use the same terminology, references, and procedures to describe details such as pores and linear discontinuities in fingerprint scans, for example.
About the Author

Gail Overton
Senior Editor (2004-2020)
Gail has more than 30 years of engineering, marketing, product management, and editorial experience in the photonics and optical communications industry. Before joining the staff at Laser Focus World in 2004, she held many product management and product marketing roles in the fiber-optics industry, most notably at Hughes (El Segundo, CA), GTE Labs (Waltham, MA), Corning (Corning, NY), Photon Kinetics (Beaverton, OR), and Newport Corporation (Irvine, CA). During her marketing career, Gail published articles in WDM Solutions and Sensors magazine and traveled internationally to conduct product and sales training. Gail received her BS degree in physics, with an emphasis in optics, from San Diego State University in San Diego, CA in May 1986.