Software simulates laser-based measurements for flow cytometry
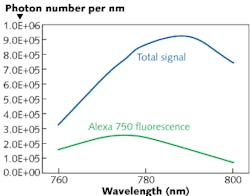
Researchers at Simphotek Inc. (Newark, NJ) have developed a simulation tool that uncovers hidden mechanisms of potential false readings in flow cytometry. Flow cytometry is used extensively in biology and medicine, where a group of cells labeled with a fluorescent probe molecule or dye is focused into a single cell stream passing through a laser light source. The fluorescent light is filtered and sampled by an array of detectors. Traditionally, one light source and one type of probe molecule/dye have been used, but additional information can be obtained if multiple lasers and multiple probes fluorescing at different wavelengths are used. By using its SimphoSOFT software, Simphotek’s R&D team recently discovered that significant false signals may occur in multiwavelength and multiprobe experiments not only due to fluorescence overlap, but also the often unanticipated phosphorescence overlap.
Phosphorescence emission generally has not been considered problematic in flow cytometry. However, numerical simulations performed by the group show that problematic or false signals from phosphorescence can range from 40% to greater than 500% of the correct (fluorescence) signal, leading to possible misinterpretations of biological results—for example, cancer cells not present when, in fact, many are. In some cases, measuring the signals in wavelength or detector channels before and after each probe is used may mitigate or reduce spectral overlap; however, false results may be unaccounted for due to limited detector sensitivity. SimphoSOFT can select a particular combination of probe molecules ahead of measurements by determining if the probe molecules may be problematic, and can correct and/or check post-experiment measurements by calculating and removing any undetected false signals. In addition to emission intensity, the group’s software also calculates photobleaching, singlet oxygen formation, energy transfer, and upconversion in multiple fluorescent probes used in biology and medicine. Contact Mary Potasek at [email protected].
About the Author
John Wallace
Senior Technical Editor (1998-2022)
John Wallace was with Laser Focus World for nearly 25 years, retiring in late June 2022. He obtained a bachelor's degree in mechanical engineering and physics at Rutgers University and a master's in optical engineering at the University of Rochester. Before becoming an editor, John worked as an engineer at RCA, Exxon, Eastman Kodak, and GCA Corporation.